236 return error(
"parameter \"files_genotyper_format\" is missing\n");
239 return error(
"parameter \"files_genotyper_logtime\" is missing\n");
243 vector<unsigned int> trait_index;
247 for(
unsigned int i = 0; i < traits.size(); i++) {
252 return error(
"Genotyper::cannot find trait \"%s\" among successfully initialized traits.\n", traits[i].c_str());
255 return error(
"Genotyper::cannot add the non-mappable trait \"%s\" to the joint genotype output file\n", traits[i].c_str());
257 trait_index.push_back(Tproto->
get_index());
264 if(format.find_last_of(
"s") == format.size()-1)
265 format = format.substr(0, format.size()-1);
267 if( format !=
"genotype" && format !=
"snp_genotype" && format !=
"snp_id")
268 return error(
"Genotyper::file format \"%s\" is not a valid option. Check the manual.\n", format.c_str());
273 for(
unsigned int i = 0; i < traits.size(); i++) {
279 if( !is2all && format.find(
"snp") != string::npos) {
280 return error(
"Genotyper:: not all traits have di-allelic loci as requested by the output format \"%s\"\n", format.c_str());
294 if(doTrimFixedLoci) {
297 warning(
"Genotyper::option \"files_genotyper_trim_maf\" is set but loci are not di-allelic. Not trimming.\n");
298 doTrimFixedLoci =
false;
301 if(format ==
"genotype") {
302 warning(
"Genotyper::option \"files_genotyper_trim_maf\" is set but data recorded is allele values (format option \"genotype\"). Not trimming.\n");
303 doTrimFixedLoci =
false;
309 warning(
"Genotyper::value %f for option \"files_genotyper_trim_maf\" cannot be used for trimming di-allelic loci.\n",trim_maf);
310 doTrimFixedLoci =
false;
333 if (format==
"genotype")
335 else if (format==
"snp_genotype")
337 else if (format==
"snp_id")
virtual void set(bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path, LCE *event)
Definition: filehandler.h:275
void set_extension(const char *ext)
Definition: filehandler.h:148
virtual void set_multi(bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path)
Definition: filehandler.h:200
FileHandler to write joint genotype files for multiple mappable traits.
Definition: servicenotifiers.h:109
void set_trimFixedLoci(bool test, double maf)
Definition: servicenotifiers.h:137
void setTraits(vector< trait_t > &traits)
Definition: servicenotifiers.cc:384
void setFormat(string format)
Definition: servicenotifiers.h:131
void setTraitIndex(vector< unsigned int > &indices)
Definition: servicenotifiers.cc:391
void set_isDiallelic(bool test)
Definition: servicenotifiers.h:135
TraitPrototype * getTraitPrototype(trait_t type)
Accessor to a TraitPrototype.
Definition: indfactory.cc:139
Metapop * _popPtr
The ptr to the current Metapop.
Definition: lifecycleevent.h:80
This structure stores one parameter, its definition and its string argument.
Definition: param.h:53
double getValue()
Returns the argument value according to its type.
Definition: param.cc:367
bool isMatrix()
Checks if the argument is of matrix type.
Definition: param.h:171
string getArg()
Definition: param.h:137
void getMatrix(TMatrix *mat)
Sets the matrix from the argument string if the parameter is set and of matrix type.
Definition: param.cc:377
vector< string > getMultiArgs()
Definition: param.cc:119
bool isSet()
Definition: param.h:139
virtual double get_parameter_value(std::string name)
Param value getter.
Definition: simcomponent.h:142
virtual Param * get_parameter(std::string name)
Param getter.
Definition: simcomponent.h:138
A class to handle matrix in params, coerces matrix into a vector of same total size.
Definition: tmatrix.h:49
TTrait setter.
Definition: ttrait.h:130
virtual int get_allele_number()=0
Returns the number of allele per locus.
virtual int get_index()
Index getter.
Definition: ttrait.h:151
virtual bool is_mappable()=0
Checks if the trait is mappable, i.e., if the loci can be placed on a genetic map.
int error(const char *str,...)
Definition: output.cc:78
void warning(const char *str,...)
Definition: output.cc:57
 Inheritance diagram for LCE_FileServicesNotifier:
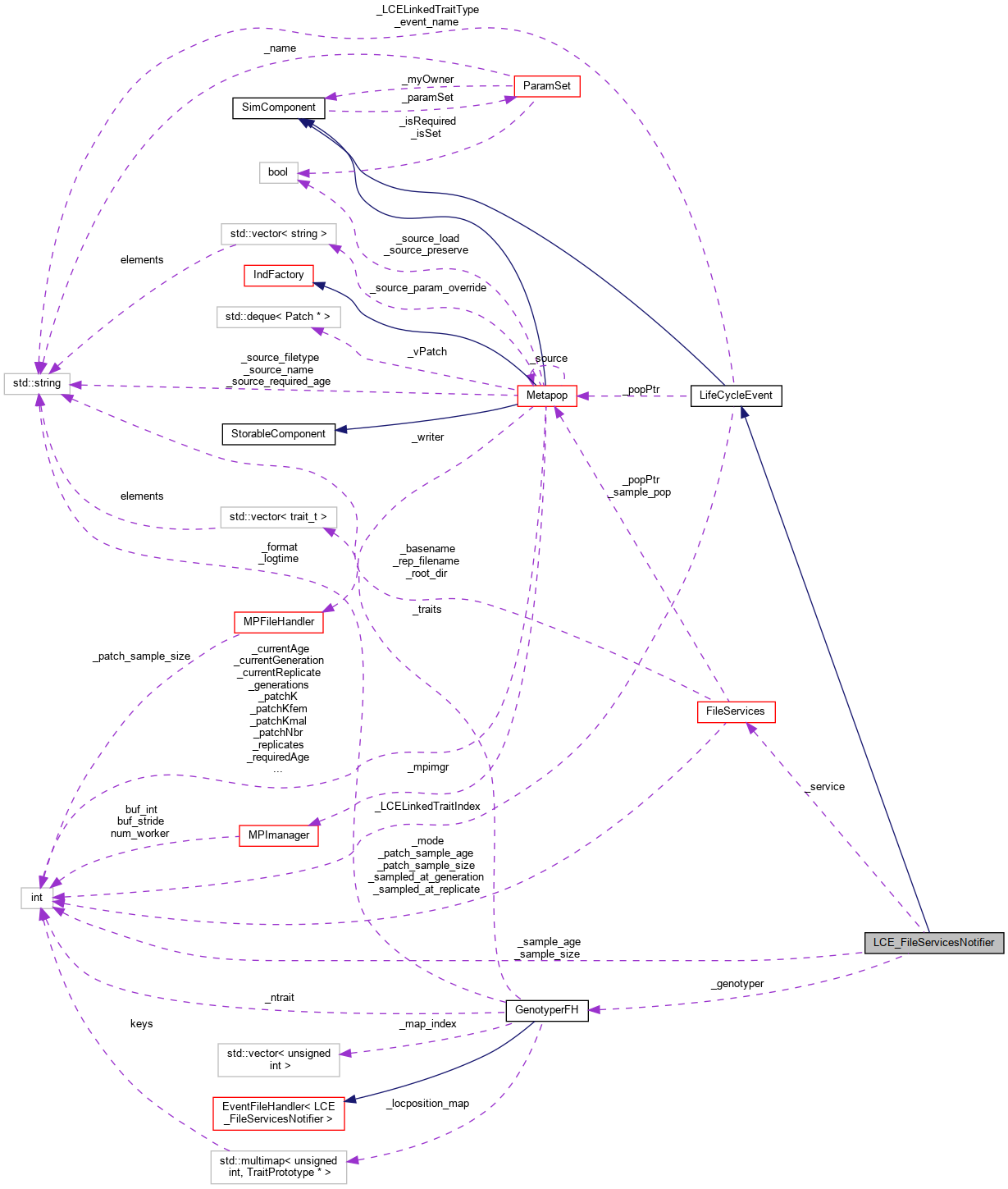
Inheritance diagram for LCE_FileServicesNotifier: Collaboration diagram for LCE_FileServicesNotifier:
Collaboration diagram for LCE_FileServicesNotifier: Public Member Functions inherited from LifeCycleEvent
Public Member Functions inherited from LifeCycleEvent Public Member Functions inherited from SimComponent
Public Member Functions inherited from SimComponent Protected Attributes inherited from LifeCycleEvent
Protected Attributes inherited from LifeCycleEvent Protected Attributes inherited from SimComponent
Protected Attributes inherited from SimComponent Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function:


 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
