TTNOhtaStats Class Reference
#include <ttneutralgenes.h>
 Inheritance diagram for TTNOhtaStats:
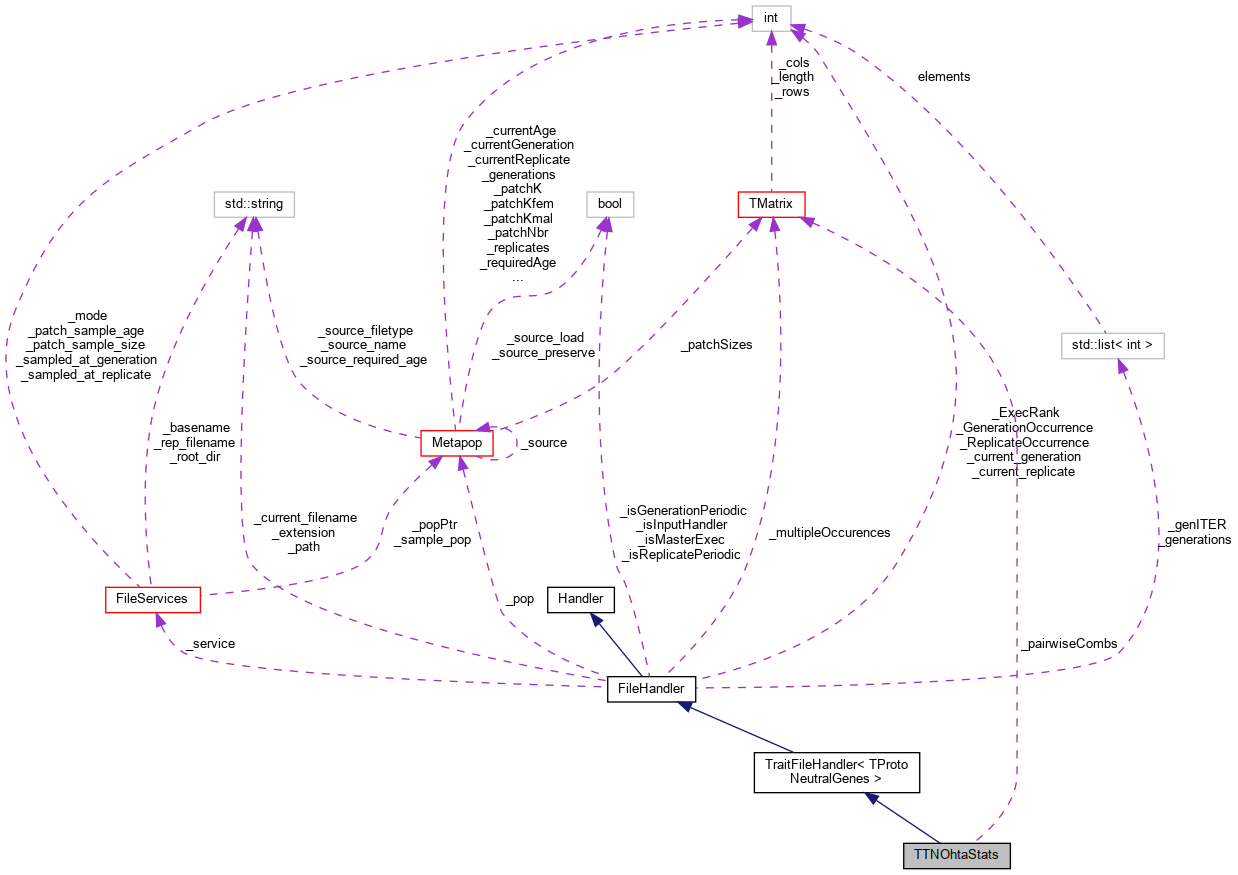
Inheritance diagram for TTNOhtaStats: Collaboration diagram for TTNOhtaStats:
Collaboration diagram for TTNOhtaStats:Public Member Functions | |
| TTNOhtaStats (TProtoNeutralGenes *T) | |
| virtual | ~TTNOhtaStats () |
| virtual void | FHwrite () |
| virtual void | FHread (string &filename) |
 Public Member Functions inherited from TraitFileHandler< TProtoNeutralGenes > Public Member Functions inherited from TraitFileHandler< TProtoNeutralGenes > | |
| TraitFileHandler (TProtoNeutralGenes *trait_proto, const char *ext) | |
| virtual | ~TraitFileHandler () |
| virtual void | FHread (string &filename)=0 |
| virtual void | set (bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path, TProtoNeutralGenes *trait_proto) |
| virtual void | set_multi (bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path, TProtoNeutralGenes *trait_proto) |
 Public Member Functions inherited from FileHandler Public Member Functions inherited from FileHandler | |
| FileHandler (const char *ext) | |
| virtual | ~FileHandler () |
| virtual void | init () |
| Called by notifier during simulation setup, performs file checking. More... | |
| virtual vector< string > | ifExist () |
| Checks if any file associated with the current file name already exists on disk. More... | |
| virtual void | set (bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path) |
| Sets the hanlder parameters. More... | |
| virtual void | set_multi (bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path) |
| virtual void | update () |
| Updates the inner replicate and generation counters and calls FHwrite if needed by the the periodicity of the file. More... | |
| Metapop * | get_pop_ptr () |
| Returns the pointer to the current metapop through the FileServices interface. More... | |
| void | set_pop_ptr (Metapop *pop_ptr) |
| FileServices * | get_service () |
| Returns pointer to the FileServices. More... | |
| void | set_service (FileServices *srv) |
| std::string & | get_path () |
| void | set_path () |
| std::string & | get_extension () |
| void | set_extension (const char *ext) |
| std::string & | get_filename () |
| Builds and returns the current file name depending on the periodicity of the file. More... | |
| bool | get_isInputHandler () |
| void | set_isInputHandler (bool val) |
| bool | get_isReplicatePeriodic () |
| void | set_isReplicatePeriodic (bool val) |
| unsigned int | get_ReplicateOccurrence () |
| void | set_ReplicateOccurrence (unsigned int val) |
| bool | get_isGenerationPeriodic () |
| void | set_isGenerationPeriodic (bool val) |
| unsigned int | get_GenerationOccurrence () |
| void | set_GenerationOccurrence (unsigned int val) |
| unsigned int | get_ExecRank () |
| unused yet... More... | |
| void | set_ExecRank (int val) |
| TMatrix * | get_OccMatrix () |
| void | set_OccMatrix (TMatrix *occ) |
| bool | get_isMasterExec () |
| void | set_isMasterExec (bool is) |
 Public Member Functions inherited from Handler Public Member Functions inherited from Handler | |
| virtual | ~Handler () |
Private Attributes | |
| TMatrix | _pairwiseCombs |
Additional Inherited Members | |
 Protected Attributes inherited from TraitFileHandler< TProtoNeutralGenes > Protected Attributes inherited from TraitFileHandler< TProtoNeutralGenes > | |
| TProtoNeutralGenes * | _FHLinkedTrait |
| int | _FHLinkedTraitIndex |
 Protected Attributes inherited from FileHandler Protected Attributes inherited from FileHandler | |
| Metapop * | _pop |
| Pointer to the current metapop, set during initialization within the init function. More... | |
Detailed Description
Constructor & Destructor Documentation
◆ TTNOhtaStats()
|
inline |
◆ ~TTNOhtaStats()
Member Function Documentation
◆ FHread()
|
inlinevirtual |
Implements FileHandler.
◆ FHwrite()
|
virtual |
Implements TraitFileHandler< TProtoNeutralGenes >.
1798 error("TTNOhtaStats::neutral trait and population are not compatible with Ohta stats (min. 2 patches, di-allelic loci), file will not be written.\n");
1816 vector< vector<double> > rSquare = vector< vector<double> > (patchNbr, vector<double>(num_comb, 0.0));
1844 unsigned int twoLocHapMap[2][2] = {{0,1},{2,3}}; // genot: AB, Ab, aB, ab; A = 0, a = 1, B = 2, b = 3
1847 vector< double > meanGenoFreq(4,0.0); // allele counter at the two loci,0 = [0], 1 = [1], 0 = [2], 1 = [3]
1850 vector< vector< double > > genoFreq = vector< vector< double > > (patchNbr, vector< double >(4,0.0));
1851 vector< vector< double > > hapFreq = vector< vector< double > > (patchNbr, vector< double >(4,0.0));
1973 Dst[pcomb] += pow((genoFreq[patch][reverseHapMap[hap][0]] * genoFreq[patch][reverseHapMap[hap][1]]) -
1984 double denom = genoFreq[patch][0] * genoFreq[patch][1] * genoFreq[patch][2] * genoFreq[patch][3];
2010 // cout << "Exp_" << reverseHapMap[hap][0] << reverseHapMap[hap][1] << " : " << (genoFreq[patch][reverseHapMap[hap][0]] * genoFreq[patch][reverseHapMap[hap][1]]) << endl;
2018 // cout << "Exp_" << reverseHapMap[hap][0] << reverseHapMap[hap][1] << " : " << (meanGenoFreq[reverseHapMap[hap][0]] * meanGenoFreq[reverseHapMap[hap][1]]) << endl;
std::string & get_filename()
Builds and returns the current file name depending on the periodicity of the file.
Definition: filehandler.cc:150
Metapop * get_pop_ptr()
Returns the pointer to the current metapop through the FileServices interface.
Definition: filehandler.h:134
This class contains traits along with other individual information (sex, pedigree,...
Definition: individual.h:48
bool isAlive()
Checks if the population still contains at least one individual in any sex or age class.
Definition: metapop.h:308
Patch * getPatch(unsigned int i)
Patch accessor, return the ith+1 patch in the metapop.
Definition: metapop.h:256
Second class in the metapopulation design structure, between the Metapop and Individual classes.
Definition: metapop.h:431
unsigned int size(age_t AGE)
Returns the size of the container of the appropriate age class(es) for both sexes.
Definition: metapop.h:497
Individual * get(sex_t SEX, age_idx AGE, unsigned int at)
Returns a pointer to the individual sitting at the index passed.
Definition: metapop.h:533
void assign(sex_t SEX, age_idx AGE, size_t n)
Assigns a new container of given size for the sex and age class passed, sets all values to NULL.
Definition: metapop.h:560
double get(unsigned int i, unsigned int j) const
Accessor to element at row i and column j.
Definition: tmatrix.h:192
virtual unsigned int get_allele(int loc, int all) const =0
Called to read the allele identity at a locus.
int _FHLinkedTraitIndex
Definition: filehandler.h:223
TProtoNeutralGenes * _FHLinkedTrait
Definition: filehandler.h:222
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTraitIndex, _pairwiseCombs, ADLTx, Patch::assign(), TMatrix::copy(), error(), fatal(), FEM, Patch::get(), TMatrix::get(), TTrait::get_allele(), TProtoNeutralGenes::get_allele_num(), FileHandler::get_filename(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_pop_ptr(), Metapop::getPatch(), Metapop::getPatchNbr(), Individual::getTrait(), Metapop::isAlive(), MAL, message(), nChooseKVec(), TMatrix::nrows(), and Patch::size().
Member Data Documentation
◆ _pairwiseCombs
The documentation for this class was generated from the following files:
 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
