|
| | TTQuanti_diallelic_no_pleio () |
| |
| | TTQuanti_diallelic_no_pleio (const TTQuanti_diallelic_no_pleio &TT) |
| |
| virtual | ~TTQuanti_diallelic_no_pleio () |
| |
| virtual void | init_sequence () |
| |
| virtual TTQuanti_diallelic_no_pleio * | clone () |
| |
| virtual void | show_up () |
| |
| virtual double | get_additive_genotype (const unsigned int trait) const |
| |
| virtual double | get_dominant_genotype (const unsigned int trait) const |
| |
| virtual void | copy_sequence_block (sex_t SEX, unsigned int strand, unsigned int from_locus, unsigned int to_locus, const TTQuanti *parent) |
| |
| virtual void | copy_sequence_1locus (sex_t SEX, unsigned int strand, unsigned int at, const TTQuanti *parent) |
| |
| | TTQuanti_diallelic () |
| |
| | TTQuanti_diallelic (const TTQuanti &T) |
| |
| virtual | ~TTQuanti_diallelic () |
| |
| virtual bool | get_allele_bit (unsigned int position, unsigned int allele) const |
| |
| virtual void | set_allele_bit (unsigned int position, unsigned int allele, bool value) |
| |
| void | inherit_free (const TTrait *mother, const TTrait *father) |
| |
| virtual void | reset () |
| |
| virtual void | init () |
| |
| virtual void | set_sequence (void **seq) |
| |
| virtual void ** | get_sequence () const |
| |
| virtual unsigned int | get_allele (int loc, int all) const |
| |
| virtual double | get_allele_value (int locus, int allele) const |
| |
| virtual void | set_allele_value (unsigned int locus, unsigned int allele, double value) |
| |
| virtual TTQuanti_diallelic & | operator= (const TTrait &T) |
| |
| virtual bool | operator== (const TTrait &T) |
| |
| virtual bool | operator!= (const TTrait &T) |
| |
| virtual void | store_data (BinaryStorageBuffer *saver) |
| |
| virtual bool | retrieve_data (BinaryStorageBuffer *reader) |
| |
| virtual double | get_full_genotype (unsigned int trait) |
| |
| virtual void | set_allele (int locus, int allele, double value) |
| |
| virtual void | mutate_add (unsigned int position, unsigned int allele, double value) |
| |
| virtual void | mutate_inplace (unsigned int position, unsigned int allele, double value) |
| |
| | TTQuanti () |
| |
| | TTQuanti (const TTQuanti &T) |
| |
| virtual | ~TTQuanti () |
| |
| virtual trait_t | get_type () const |
| |
| virtual void | mutate () |
| |
| virtual void | inherit (const TTrait *mother, const TTrait *father) |
| |
| virtual void * | set_trait (void *value) |
| |
| virtual void | set_value () |
| |
| virtual void * | getValue () const |
| |
| void | set_proto (TProtoQuanti *proto) |
| |
| TProtoQuanti * | get_proto () |
| |
| double | get_phenotype (unsigned int trai) |
| |
| void | set_phenotype (unsigned int trait, double value) |
| |
| void | reset_phenotype_to_genotypic_value () |
| |
| virtual | ~TTrait () |
| |
| virtual | ~StorableComponent () |
| |
TTQuanti_diallelic_no_pleio : single or multiple non-pleiotropic traits, di-allelic.
 Inheritance diagram for TTQuanti_diallelic_no_pleio:
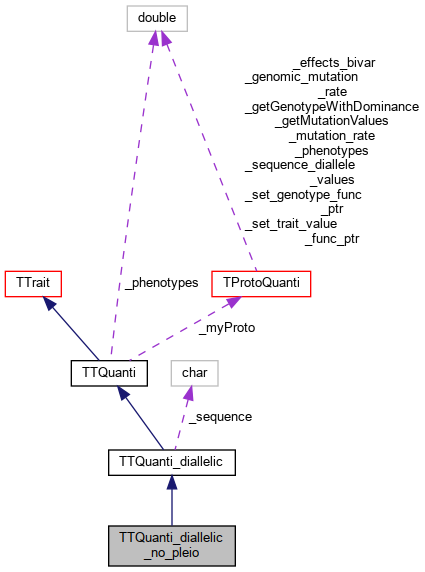
Inheritance diagram for TTQuanti_diallelic_no_pleio: Collaboration diagram for TTQuanti_diallelic_no_pleio:
Collaboration diagram for TTQuanti_diallelic_no_pleio: Public Member Functions inherited from TTQuanti_diallelic
Public Member Functions inherited from TTQuanti_diallelic Public Member Functions inherited from TTQuanti
Public Member Functions inherited from TTQuanti Public Member Functions inherited from TTrait
Public Member Functions inherited from TTrait Public Member Functions inherited from StorableComponent
Public Member Functions inherited from StorableComponent Protected Attributes inherited from TTQuanti_diallelic
Protected Attributes inherited from TTQuanti_diallelic Protected Attributes inherited from TTQuanti
Protected Attributes inherited from TTQuanti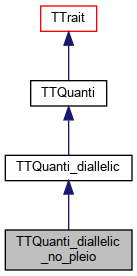

 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
