#include <ttquanti.h>
 Inheritance diagram for TProtoQuanti:
Inheritance diagram for TProtoQuanti: Collaboration diagram for TProtoQuanti:
Collaboration diagram for TProtoQuanti:Public Member Functions | |
| TProtoQuanti () | |
| TProtoQuanti (const TProtoQuanti &T) | |
| virtual | ~TProtoQuanti () |
| unsigned int | get_num_traits () |
| unsigned int | get_num_locus () |
| unsigned int | get_num_locus (unsigned int trait) |
| unsigned int | get_pleiotropy_type () |
| unsigned int | get_seq_length () |
| size_t | get_size_locus_type () |
| size_t | get_locus_byte_size () |
| vector< double > | get_env_var () |
| vector< double > | get_heritability () |
| unsigned int | get_h2_setTime () |
| bool | get_h2_isBroad () |
| unsigned int | get_allele_model () |
| const TMatrix & | get_diallele_values () |
| double | get_diallele_value (unsigned int locus, unsigned int allele) |
| double | get_seq_diallele_value (unsigned int position, unsigned int allele) |
| bool | has_equal_domCoeff () |
| bool | has_equal_effects () |
| double | get_equal_val_0 () |
| double | get_equal_val_1 () |
| double | get_equal_val_diff () |
| const bitstring & | get_trait_mask (unsigned int trait) const |
| double | get_init_value (unsigned int i) |
| double | get_init_variance (unsigned int i) |
| unsigned int | get_doInitMutation () |
| TTQuantiSH * | get_stater () |
| const TMatrix & | get_pleio_matrix () |
| unsigned int | get_locus_seq_pos (unsigned int loc, unsigned int trait) |
| unsigned int | get_locus_ID (unsigned int locus, unsigned int trait) |
| unsigned int | get_locus_PD (unsigned int locus) |
| unsigned int | get_locus_start_pos (unsigned int locus) |
| unsigned int | get_sequence_block_size (unsigned int from, unsigned int to) |
| void | set_eVarianceSD (unsigned int trait, double SD) |
| void | set_init_values (const double *values, unsigned int nval) |
| void | set_trait_value_func_ptr (bool withVe) |
| double | set_trait_value_VE (const TTQuanti *ind, const unsigned int trait) |
| double | set_trait_value_noVE (const TTQuanti *ind, const unsigned int trait) |
| double | set_genotype_value_additive (const TTQuanti *ind, const unsigned int trait) |
| double | set_genotype_value_dominance (const TTQuanti *ind, const unsigned int trait) |
| int | get_allele_position (const unsigned int locus, const unsigned int trait) |
| double | get_allele_value (const TTQuanti *ind, const unsigned int allele, const unsigned int locus, const unsigned int trait) |
| double | get_genotype_with_dominance (const double a1, const double a2, const unsigned int locus, const unsigned int trait) |
| double | get_genotype_dominance_h (double a1, double a2, double h) |
| double | get_genotype_dominance_k (double a1, double a2, double k) |
| unsigned int | get_dominance_model () |
| double | get_dominance (unsigned int locus, unsigned int trait) |
| const TMatrix & | get_dominance_effects () |
| double | get_trait_mutation_variance (unsigned int trait) |
| double | get_genotypic_value (const TTQuanti *ind, const unsigned int trait) |
| double | get_phenotypic_value (const TTQuanti *ind, const unsigned int trait) |
| bool | setHeritabilityParams () |
| bool | setInitialValuesParams () |
| bool | setGeneticMapParams () |
| unsigned int | setAlleleModel () |
| bool | setMutationModel_no_pleio () |
| bool | setMutationModel_full_pleio () |
| bool | setMutationModel_var_pleio () |
| void | setTraitAndLocusTables_full_pleio () |
| void | setTraitAndLocusTables_no_pleio (TMatrix &mat) |
| bool | setMutationCorrelation () |
| bool | setDominanceParameters () |
| bool | setDiallelicMutationModel () |
| bool | setMutationSigmaFromQuantiMutationVariance () |
| bool | setMutationSigmaFromQuantiMutationVariance_no_pleio () |
| bool | readMatrixFromQuantiMutationMatrix (vector< vector< double >> &varmat) |
| bool | setContinuousMutationModel_no_pleio () |
| bool | setContinuousMutationModel_full_pleio () |
| bool | setContinuousMutationModel_var_pleio () |
| void | allocate_gsl_mutation_matrix_space (unsigned int num_locus) |
| void | deallocate_gsl_mutation_matrix_space () |
| gsl_matrix * | set_gsl_mutation_matrix_from_sigma (unsigned int loc, unsigned int pleio_deg) |
| gsl_matrix * | set_gsl_mutation_matrix (unsigned int pleio_deg, const vector< double > &varcov) |
| void | set_mutation_matrix_decomposition (unsigned int loc, unsigned int pleio_deg) |
| double * | getMutationEffectMultivariateGaussian (unsigned int loc) |
| double * | getMutationEffectBivariateGaussian (unsigned int loc) |
| double * | getMutationEffectUnivariateGaussian (unsigned int loc) |
| double * | getMutationEffectMultivariateGaussianLocSpec (unsigned int loc) |
| double * | getMutationEffectBivariateGaussianLocSpec (unsigned int loc) |
| double * | getMutationEffectUnivariateGaussianLocSpec (unsigned int loc) |
| double * | getMutationEffectBivariateDiallelic (unsigned int loc) |
| double * | getMutationEffects (unsigned int loc) |
| double * | getMutationEffectsVarPleio (unsigned int loc) |
| void | inherit (sex_t SEX, TTQuanti *ind, const TTQuanti *parent) |
| void | inherit_free (sex_t SEX, TTQuanti *ind, const TTQuanti *parent) |
| void | inherit_low (sex_t SEX, TTQuanti *ind, const TTQuanti *parent) |
| unsigned int | get_num_mutations () |
| void | mutate (TTQuanti *ind) |
| void | mutate_nill (TTQuanti *ind) |
| void | mutate_full_pleio (TTQuanti *ind) |
| void | mutate_var_pleio (TTQuanti *ind) |
| void | mutate_no_pleio (TTQuanti *ind) |
| void | mutate_inplace_full_pleio (TTQuanti *ind) |
| void | mutate_inplace_var_pleio (TTQuanti *ind) |
| void | mutate_inplace_no_pleio (TTQuanti *ind) |
| void | mutate_diallelic_pleio (TTQuanti *ind) |
| void | mutate_diallelic_var_pleio (TTQuanti *ind) |
| void | mutate_diallelic_no_pleio (TTQuanti *ind) |
| bool | setEpistasisParameters () |
| unsigned int | get_num_epi_coefs () |
| bool | do_epistasis () |
| const TMatrix & | get_epi_coefs () const |
| const TMatrix & | get_epi_coef_index () const |
TraitPrototype implementations | |
| virtual void | reset () |
| virtual TTQuanti * | hatch () |
| virtual TProtoQuanti * | clone () |
| virtual trait_t | get_type () const |
| virtual int | get_phenotype_dimension () |
| Returns the dimension of the phenotype of the trait (size of the array accessed with TTrait::getValue() More... | |
| virtual int | get_allele_number () |
| Returns the number of allele per locus. More... | |
| virtual int | get_locus_number () |
| Returns the number of locus. More... | |
StorableComponent implementation | |
| virtual void | store_data (BinaryStorageBuffer *saver) |
| virtual bool | retrieve_data (BinaryStorageBuffer *reader) |
SimComponent implementation | |
| virtual bool | setParameters () |
| virtual void | loadFileServices (FileServices *loader) |
| virtual void | loadStatServices (StatServices *loader) |
| virtual bool | resetParameterFromSource (std::string param, SimComponent *cmpt) |
 Public Member Functions inherited from TTProtoWithMap Public Member Functions inherited from TTProtoWithMap | |
| TTProtoWithMap () | |
| TTProtoWithMap (const TTProtoWithMap &TP) | |
| virtual | ~TTProtoWithMap () |
| void | setMapIndex (unsigned int idx) |
| unsigned int | getMapIndex () |
| double | getRecombRate () const |
| bool | setGeneticMapParameters (string prefix, unsigned int numLoci=0) |
| void | addGeneticMapParameters (string prefix) |
| bool | setRecombinationMapRandom () |
| bool | setRecombinationMapNonRandom (vector< vector< double > > &lociPositions) |
| bool | setRecombinationMapFixed () |
| bool | setNumLociPerChromosome (string param_name) |
| void | reset_recombination_pointers () |
| void | registerGeneticMap () |
| void | unregisterFromGeneticMap () |
| bool | areGeneticMapParamSet (string prefix) |
| bool | isRecombinationFree (string prefix) |
| void | recordRandomMap () |
| virtual bool | is_mappable () |
| Checks if the trait is mappable, i.e., if the loci can be placed on a genetic map. More... | |
| virtual bool | is_mapped () |
| Checks if the trait's loci are placed on a genetic map. More... | |
| virtual vector< unsigned int > | get_locus_map_positions () |
| Returns the map positions of the loci in vector. More... | |
 Public Member Functions inherited from TraitPrototype Public Member Functions inherited from TraitPrototype | |
| virtual void | set_index (int idx) |
| Sets the traits index. More... | |
| virtual int | get_index () |
| Index getter. More... | |
 Public Member Functions inherited from StorableComponent Public Member Functions inherited from StorableComponent | |
| virtual | ~StorableComponent () |
 Public Member Functions inherited from SimComponent Public Member Functions inherited from SimComponent | |
| SimComponent () | |
| virtual | ~SimComponent () |
| virtual void | loadUpdaters (UpdaterServices *loader) |
| Loads the parameters and component updater onto the updater manager. More... | |
| virtual void | set_paramset (ParamSet *paramset) |
| Sets the ParamSet member. More... | |
| virtual void | set_paramset (std::string name, bool required, SimComponent *owner) |
| Sets a new ParamSet and name it. More... | |
| virtual void | set_paramsetFromCopy (const ParamSet &PSet) |
| Reset the set of parameters from a another set. More... | |
| virtual ParamSet * | get_paramset () |
| ParamSet accessor. More... | |
| virtual void | add_parameter (Param *param) |
| Interface to add a parameter to the set. More... | |
| virtual void | add_parameter (std::string Name, param_t Type, bool isRequired, bool isBounded, double low_bnd, double up_bnd) |
| Interface to add a parameter to the set. More... | |
| virtual void | add_parameter (std::string Name, param_t Type, bool isRequired, bool isBounded, double low_bnd, double up_bnd, ParamUpdaterBase *updater) |
| Interface to add a parameter and its updater to the set. More... | |
| virtual Param * | get_parameter (std::string name) |
| Param getter. More... | |
| virtual double | get_parameter_value (std::string name) |
| Param value getter. More... | |
| virtual string | get_name () |
| Returnd the name of the ParamSet, i.e. More... | |
| virtual bool | has_parameter (std::string name) |
| Param getter. More... | |
Protected Attributes | |
| unsigned int | _num_locus |
| Total number of loci, for all traits. More... | |
| unsigned int | _2L |
| Diploid locus size, to save on useless operations during mutation. More... | |
| unsigned int | _num_traits |
| Number of traits. More... | |
| unsigned int | _seq_length |
| Total number of allelic values stored in individual sequences, no trait x no locus. More... | |
| unsigned int | _allele_model |
| string | _diallele_datatype |
| double | _mutation_rate |
| double | _genomic_mutation_rate |
| unsigned int | _pleio_type |
 Protected Attributes inherited from TTProtoWithMap Protected Attributes inherited from TTProtoWithMap | |
| unsigned int | _mapIndex |
| double | _totRecombEventsMean |
| double | _recombRate |
| double | _mapResolution |
| unsigned int | _numChromosome |
| unsigned int | _numLoci |
| double * | _recombRatePerChrmsm |
| unsigned int * | _numLociPerChrmsm |
| unsigned int * | _chrsmLength |
| unsigned int * | _lociMapPositions |
 Protected Attributes inherited from TraitPrototype Protected Attributes inherited from TraitPrototype | |
| int | _index |
| The trait index in the Individual traits table. More... | |
 Protected Attributes inherited from SimComponent Protected Attributes inherited from SimComponent | |
| ParamSet * | _paramSet |
| The parameters container. More... | |
Private Attributes | |
| TMatrix | _allele_value |
| double * | _sequence_diallele_values [2] |
| bool | _equal_effects |
| double | _equal_val_0 |
| double | _equal_val_1 |
| double | _equal_val_diff |
| vector< bitstring > | _trait_masks |
| TMatrix | _init_value |
| TMatrix | _init_variance |
| TMatrix | _mutation_correlation |
| TMatrix | _mutation_sigma |
| gsl_matrix ** | _gsl_mutation_matrix |
| gsl_matrix ** | _evect |
| gsl_vector ** | _eval |
| gsl_vector ** | _effects_multivar |
| gsl_vector ** | _ws |
| double | _effects_bivar [2] |
| unsigned int | _doInitMutation |
| bool | _mutationVarianceIsLocusSpecific |
| bool | _mutationEffectIsFixedDiAllele |
| unsigned int | _dominance_model |
| TMatrix | _dominance_effects |
| bool | _equal_dom_coeff |
| bool * | _all_chooser |
| size_t | _locusByteSize |
| size_t | _sizeofLocusType |
| vector< double > | _eVariance |
| vector< double > | _h2 |
| unsigned int | _h2_setTime |
| bool | _h2_isBroad |
| TMatrix | _pleio_matx |
| Pleiotropy matrix provided in input (num locu X num trait). More... | |
| vector< vector< unsigned int > > | _trait_table |
| Trait table, (num trait X (variable length/trait)), holds, for each trait, the array position of causative alleles in the sequence. More... | |
| vector< vector< unsigned int > > | _trait_locus_table |
| Table storing the locus id of each locus affecting each trait (num trait X (variable length/trait)). More... | |
| vector< vector< unsigned int > > | _locus_table |
| Locus table, num_locus x 2, first column holds the start position of the alleles of each locus in the sequence, second column counts the number of alleles = pleiotropic degree. More... | |
| bool | _epistasis |
| unsigned int | _num_epi_coefs |
| TMatrix | _epistatic_coefs_matrix |
| TMatrix | _epistatic_coefs_indices |
| void(TProtoQuanti::* | _inherit_fun_ptr )(sex_t, TTQuanti *, const TTQuanti *) |
| Pointer to inheritance functions: either inherit_free() (r=0.5), or inherit_low() (r<0.5). More... | |
| void(TProtoQuanti::* | _mutation_func_ptr )(TTQuanti *) |
| Pointer to mutation function, which depends on allele on model (HC, noHC, diallelic) More... | |
| double *(TProtoQuanti::* | _getMutationValues )(unsigned int) |
| Pointer to mutation allele value function, which depends on allele model and number of traits affected. More... | |
| vector< double *(TProtoQuanti::*)(unsigned int) > | _getMutationValuesVarPleio |
| Collection of pointers to mutation functions, which generate allele values in dependence of pleiotropic degree. More... | |
| double(TProtoQuanti::* | _getGenotypeWithDominance )(double, double, double) |
| Pointer to either dominance_h() or dominance_k() function computing the genotypic value with dominance. More... | |
| double(TProtoQuanti::* | _set_trait_value_func_ptr )(const TTQuanti *, const unsigned int) |
| Pointer to either set_trait_value_VE() or set_trait_value_noVE() to compute phenotypic values. More... | |
| double(TProtoQuanti::* | _set_genotype_func_ptr )(const TTQuanti *, const unsigned int) |
| Pointer to functions get_genotype_value_additive() or get_genotype_value_dominance() computing the genotypic value of a trait as function of allele effect. More... | |
| TTQuantiSH * | _stats |
| TTQuantiFH * | _writer |
| TTQuantiFH * | _reader |
| TTQFreqExtractor * | _freqExtractor |
| TTQOhtaStats * | _ohtaStats |
Friends | |
| class | TTQuanti |
| class | TTQuanti_continuous |
| class | TTQuanti_continuous_full_pleio |
| class | TTQuanti_continuous_var_pleio |
| class | TTQuanti_continuous_no_pleio |
| class | TTQuanti_continuous_single |
Additional Inherited Members | |
 Static Public Member Functions inherited from TTProtoWithMap Static Public Member Functions inherited from TTProtoWithMap | |
| static GeneticMap & | getGeneticMapRef () |
| static void | recombine (unsigned long indID) |
 Static Public Attributes inherited from TTProtoWithMap Static Public Attributes inherited from TTProtoWithMap | |
| static GeneticMap | _map |
Detailed Description
Constructor & Destructor Documentation
◆ TProtoQuanti() [1/2]
| TProtoQuanti::TProtoQuanti | ( | ) |
References _equal_effects, _equal_val_0, _equal_val_1, _equal_val_diff, _sequence_diallele_values, SimComponent::add_parameter(), TTProtoWithMap::addGeneticMapParameters(), BOOL, DBL, INT, MAT, SimComponent::set_paramset(), and STR.
Referenced by clone().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ TProtoQuanti() [2/2]
| TProtoQuanti::TProtoQuanti | ( | const TProtoQuanti & | T | ) |
References _epistatic_coefs_indices, _epistatic_coefs_matrix, _equal_effects, _equal_val_0, _equal_val_1, _equal_val_diff, _locusByteSize, _num_traits, SimComponent::_paramSet, _pleio_matx, _sequence_diallele_values, and TMatrix::copy().
◆ ~TProtoQuanti()
|
virtual |
References _all_chooser, _freqExtractor, _ohtaStats, _reader, _sequence_diallele_values, _stats, _writer, and deallocate_gsl_mutation_matrix_space().
Member Function Documentation
◆ allocate_gsl_mutation_matrix_space()
| void TProtoQuanti::allocate_gsl_mutation_matrix_space | ( | unsigned int | num_locus | ) |
References _effects_multivar, _eval, _evect, _gsl_mutation_matrix, and _ws.
Referenced by setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ clone()
|
inlinevirtual |
◆ deallocate_gsl_mutation_matrix_space()
| void TProtoQuanti::deallocate_gsl_mutation_matrix_space | ( | ) |
References _effects_multivar, _eval, _evect, _gsl_mutation_matrix, _mutationVarianceIsLocusSpecific, _num_locus, and _ws.
Referenced by setParameters(), and ~TProtoQuanti().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ do_epistasis()
|
inline |
References _epistasis.
Referenced by TTQuantiSH::addQuanti(), and TTQuantiSH::addVarPerPatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_allele_model()
|
inline |
References _allele_model.
Referenced by TTQuantiFH::FHread(), TTQFreqExtractor::FHwrite(), TTQOhtaStats::FHwrite(), TTQuantiSH::getSNPalleleFreqInPatch(), TTQuantiSH::getVaWithDominance(), loadFileServices(), LCE_QuantiInit::setParameters(), TTQuantiFH::write_PLINK(), and TTQuantiFH::write_TABLE().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_allele_number()
|
inlinevirtual |
◆ get_allele_position()
| int TProtoQuanti::get_allele_position | ( | const unsigned int | locus, |
| const unsigned int | trait | ||
| ) |
References _trait_locus_table, and _trait_table.
Referenced by get_allele_value().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_allele_value()
| double TProtoQuanti::get_allele_value | ( | const TTQuanti * | ind, |
| const unsigned int | allele, | ||
| const unsigned int | locus, | ||
| const unsigned int | trait | ||
| ) |
References error(), get_allele_position(), and TTrait::get_allele_value().
◆ get_diallele_value()
|
inline |
References _allele_value, and TMatrix::get().
◆ get_diallele_values()
|
inline |
References _allele_value.
Referenced by TTQFreqExtractor::FHwrite(), TTQuantiSH::getVaNoDominance(), and TTQuantiSH::getVaWithDominance().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_doInitMutation()
|
inline |
References _doInitMutation.
Referenced by TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_diallelic_no_pleio::init_sequence(), TTQuanti_diallelic_full_pleio::init_sequence(), TTQuanti_diallelic_var_pleio::init_sequence(), TTQuanti_diallelic_bitstring_no_pleio::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio::init_sequence(), TTQuanti_diallelic_bitstring_var_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), TTQuanti_diallelic_full_pleio_epistasis::init_sequence(), TTQuanti_continuous_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::init_sequence(), and TTQuanti_diallelic_bitstring_full_pleio_epistasis::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_dominance()
|
inline |
References _dominance_effects, and TMatrix::get().
Referenced by TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype(), and get_genotype_with_dominance().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_dominance_effects()
|
inline |
References _dominance_effects.
Referenced by TTQuanti_continuous_no_pleio::get_dominant_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_dominance_model()
|
inline |
References _dominance_model.
Referenced by TTQuantiSH::addQuanti(), TTQuantiSH::addVarPerPatch(), TTQuantiFH::FHread(), LCE_QuantiModifier::setParameters(), LCE_Breed_Quanti::setParameters(), and TTQuantiFH::write_TABLE().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_env_var()
|
inline |
References _eVariance.
Referenced by TTQuantiFH::FHread(), TTQuantiSH::init(), and TTQuantiFH::write_TABLE().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_epi_coef_index()
|
inline |
References _epistatic_coefs_indices.
Referenced by TTQuanti_continuous_full_pleio_epistasis::get_epistatic_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_epistatic_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_epistatic_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_epistatic_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_epistatic_genotype(), and TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_epistatic_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_epi_coefs()
|
inline |
References _epistatic_coefs_matrix.
Referenced by TTQuanti_continuous_full_pleio_epistasis::get_epistatic_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_epistatic_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_epistatic_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_epistatic_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_epistatic_genotype(), and TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_epistatic_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_equal_val_0()
|
inline |
References _equal_val_0.
Referenced by TTQuanti_diallelic_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype_equal_effects(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), and TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_equal_val_1()
|
inline |
References _equal_val_1.
Referenced by TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), and TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_equal_val_diff()
|
inline |
References _equal_val_diff.
Referenced by TTQuanti_diallelic_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), and TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype_equal_effects().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_genotype_dominance_h()
|
inline |
◆ get_genotype_dominance_k()
|
inline |
Referenced by get_genotype_with_dominance(), and setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_genotype_with_dominance()
| double TProtoQuanti::get_genotype_with_dominance | ( | const double | a1, |
| const double | a2, | ||
| const unsigned int | locus, | ||
| const unsigned int | trait | ||
| ) |
References get_dominance(), and get_genotype_dominance_k().
Referenced by TTQuanti_continuous_full_pleio::get_dominant_genotype(), TTQuanti_continuous_var_pleio::get_dominant_genotype(), TTQuanti_continuous_single::get_dominant_genotype(), TTQuanti_diallelic_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype(), TTQuanti_continuous_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_dominant_genotype(), and TTQuantiSH::getVaWithDominance().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_genotypic_value()
|
inline |
References _set_genotype_func_ptr.
Referenced by TTQuanti_continuous::get_full_genotype(), TTQuanti_diallelic::get_full_genotype(), and TTQuanti_diallelic_bitstring::get_full_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_h2_isBroad()
|
inline |
References _h2_isBroad.
Referenced by TTQuantiSH::getVaWithDominance().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_h2_setTime()
|
inline |
References _h2_setTime.
Referenced by LCE_Breed_Quanti::setVefromVa().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_heritability()
|
inline |
References _h2.
Referenced by LCE_QuantiModifier::execute(), TTQuantiSH::init(), LCE_Breed_Quanti::NonWrightFisherPopulation(), and LCE_Breed_Quanti::WrightFisherPopulation().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_init_value()
|
inline |
References _init_value, and TMatrix::get().
Referenced by TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), and TTQuanti_continuous_no_pleio_epistasis::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_init_variance()
|
inline |
References _init_variance, and TMatrix::get().
Referenced by TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), and TTQuanti_continuous_no_pleio_epistasis::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_locus_byte_size()
|
inline |
References _locusByteSize.
Referenced by TTQuanti_continuous_full_pleio_epistasis::copy_sequence_1locus(), TTQuanti_diallelic_full_pleio_epistasis::copy_sequence_1locus(), TTQuanti_continuous_full_pleio::copy_sequence_1locus(), TTQuanti_diallelic_full_pleio::copy_sequence_1locus(), TTQuanti_continuous_full_pleio_epistasis::copy_sequence_block(), TTQuanti_diallelic_full_pleio_epistasis::copy_sequence_block(), TTQuanti_continuous_no_pleio_epistasis::copy_sequence_block(), TTQuanti_diallelic_no_pleio_epistasis::copy_sequence_block(), TTQuanti_continuous_full_pleio::copy_sequence_block(), TTQuanti_continuous_no_pleio::copy_sequence_block(), TTQuanti_diallelic_no_pleio::copy_sequence_block(), and TTQuanti_diallelic_full_pleio::copy_sequence_block().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_locus_ID()
|
inline |
References _trait_locus_table.
Referenced by TTQFreqExtractor::FHwrite(), TTQuanti_continuous_var_pleio::get_dominant_genotype(), TTQuanti_continuous_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_dominant_genotype(), TTQuantiSH::getVaWithDominance(), setDiallelicMutationModel(), TTQuanti_continuous_var_pleio::show_up(), TTQuanti_continuous_no_pleio::show_up(), TTQuanti_diallelic_no_pleio::show_up(), TTQuanti_diallelic_var_pleio::show_up(), TTQuanti_diallelic_bitstring_no_pleio::show_up(), TTQuanti_diallelic_bitstring_var_pleio::show_up(), TTQuanti_continuous_no_pleio_epistasis::show_up(), TTQuanti_diallelic_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::show_up(), TTQuantiFH::write_PLINK(), and TTQuantiFH::write_TABLE().
◆ get_locus_number()
|
inlinevirtual |
◆ get_locus_PD()
|
inline |
References _locus_table.
Referenced by TTQuanti_diallelic_bitstring_var_pleio::copy_sequence_1locus(), TTQuanti_continuous_var_pleio::copy_sequence_1locus(), TTQuanti_diallelic_var_pleio::copy_sequence_1locus(), getMutationEffectMultivariateGaussian(), getMutationEffectMultivariateGaussianLocSpec(), and TTQuanti_continuous_var_pleio::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_locus_seq_pos()
|
inline |
References _trait_table.
Referenced by TTQuantiFH::FHread(), TTQFreqExtractor::FHwrite(), TTQuanti_continuous_var_pleio::get_additive_genotype(), TTQuanti_continuous_no_pleio::get_additive_genotype(), TTQuanti_diallelic_no_pleio::get_additive_genotype(), TTQuanti_diallelic_full_pleio::get_additive_genotype(), TTQuanti_diallelic_var_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_additive_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype_equal_effects(), TTQuanti_continuous_var_pleio::get_dominant_genotype(), TTQuanti_continuous_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_dominant_genotype(), TTQuantiSH::getVaWithDominance(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_diallelic_full_pleio::init_sequence(), TTQuanti_diallelic_var_pleio::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio::init_sequence(), TTQuanti_diallelic_bitstring_var_pleio::init_sequence(), TTQuanti_diallelic_full_pleio_epistasis::init_sequence(), TTQuanti_continuous_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::init_sequence(), TTQuantiFH::print(), TTQuantiFH::print_PLINK_PED(), setDiallelicMutationModel(), TTQuanti_continuous_var_pleio::show_up(), TTQuanti_continuous_no_pleio::show_up(), TTQuanti_diallelic_no_pleio::show_up(), TTQuanti_diallelic_full_pleio::show_up(), TTQuanti_diallelic_var_pleio::show_up(), TTQuanti_diallelic_bitstring_no_pleio::show_up(), TTQuanti_diallelic_bitstring_full_pleio::show_up(), TTQuanti_diallelic_bitstring_var_pleio::show_up(), TTQuanti_diallelic_full_pleio_epistasis::show_up(), TTQuanti_continuous_no_pleio_epistasis::show_up(), TTQuanti_diallelic_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::show_up(), and TTQuanti_diallelic_bitstring_full_pleio_epistasis::show_up().
◆ get_locus_start_pos()
|
inline |
References _locus_table.
Referenced by TTQuanti_diallelic_bitstring_full_pleio::copy_sequence_1locus(), TTQuanti_diallelic_bitstring_var_pleio::copy_sequence_1locus(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::copy_sequence_1locus(), TTQuanti_continuous_var_pleio::copy_sequence_1locus(), TTQuanti_diallelic_var_pleio::copy_sequence_1locus(), TTQuanti_diallelic_bitstring_full_pleio::copy_sequence_block(), TTQuanti_diallelic_bitstring_var_pleio::copy_sequence_block(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::copy_sequence_block(), TTQuanti_continuous_var_pleio::copy_sequence_block(), TTQuanti_diallelic_var_pleio::copy_sequence_block(), and TTQuanti_continuous_var_pleio::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_num_epi_coefs()
|
inline |
References _num_epi_coefs.
◆ get_num_locus() [1/2]
|
inline |
References _num_locus.
Referenced by TTQuantiFH::FHread(), TTQFreqExtractor::FHwrite(), TTQOhtaStats::FHwrite(), TTQuanti_continuous_full_pleio::get_additive_genotype(), TTQuanti_continuous_var_pleio::get_additive_genotype(), TTQuanti_continuous_no_pleio::get_additive_genotype(), TTQuanti_continuous_single::get_additive_genotype(), TTQuanti_diallelic_no_pleio::get_additive_genotype(), TTQuanti_diallelic_full_pleio::get_additive_genotype(), TTQuanti_diallelic_var_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), TTQuanti_continuous_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_additive_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype_equal_effects(), TTQuanti_continuous_full_pleio::get_dominant_genotype(), TTQuanti_continuous_var_pleio::get_dominant_genotype(), TTQuanti_continuous_no_pleio::get_dominant_genotype(), TTQuanti_continuous_single::get_dominant_genotype(), TTQuanti_diallelic_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype(), TTQuanti_continuous_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_continuous_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_dominant_genotype(), TTQuantiSH::getVaWithDominance(), TTQuantiSH::init(), TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_diallelic_no_pleio::init_sequence(), TTQuanti_diallelic_full_pleio::init_sequence(), TTQuanti_diallelic_var_pleio::init_sequence(), TTQuanti_diallelic_bitstring_no_pleio::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio::init_sequence(), TTQuanti_diallelic_bitstring_var_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), TTQuanti_diallelic_full_pleio_epistasis::init_sequence(), TTQuanti_continuous_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::init_sequence(), TTQuanti_continuous::operator==(), TTQuanti_diallelic::operator==(), TTQuanti_diallelic_bitstring::operator==(), TTQuantiFH::print(), TTQuantiFH::print_PLINK_PED(), setDiallelicMutationModel(), LCE_QuantiInit::setParameters(), TTQuanti_continuous_full_pleio::show_up(), TTQuanti_continuous_var_pleio::show_up(), TTQuanti_continuous_no_pleio::show_up(), TTQuanti_diallelic_no_pleio::show_up(), TTQuanti_diallelic_full_pleio::show_up(), TTQuanti_diallelic_var_pleio::show_up(), TTQuanti_diallelic_bitstring_no_pleio::show_up(), TTQuanti_diallelic_bitstring_full_pleio::show_up(), TTQuanti_diallelic_bitstring_var_pleio::show_up(), TTQuanti_continuous_full_pleio_epistasis::show_up(), TTQuanti_diallelic_full_pleio_epistasis::show_up(), TTQuanti_continuous_no_pleio_epistasis::show_up(), TTQuanti_diallelic_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::show_up(), TTQuantiFH::write_PLINK(), and TTQuantiFH::write_TABLE().
◆ get_num_locus() [2/2]
|
inline |
References _trait_table.
◆ get_num_mutations()
|
inline |
References _2L, _mutation_rate, and RAND::Binomial().
Referenced by mutate_diallelic_no_pleio(), mutate_diallelic_pleio(), mutate_diallelic_var_pleio(), mutate_full_pleio(), mutate_inplace_full_pleio(), mutate_inplace_no_pleio(), mutate_inplace_var_pleio(), mutate_no_pleio(), and mutate_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_num_traits()
|
inline |
References _num_traits.
Referenced by LCE_PhenotypeExpression::check_g_index_matrix(), TTQuanti_diallelic_bitstring_full_pleio::copy_sequence_1locus(), TTQuanti_continuous_full_pleio_epistasis::copy_sequence_1locus(), TTQuanti_diallelic_full_pleio_epistasis::copy_sequence_1locus(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::copy_sequence_1locus(), TTQuanti_continuous_full_pleio::copy_sequence_1locus(), TTQuanti_diallelic_full_pleio::copy_sequence_1locus(), TTQuanti_diallelic_bitstring_full_pleio::copy_sequence_block(), TTQuanti_continuous_full_pleio_epistasis::copy_sequence_block(), TTQuanti_diallelic_full_pleio_epistasis::copy_sequence_block(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::copy_sequence_block(), TTQuanti_continuous_full_pleio::copy_sequence_block(), TTQuanti_diallelic_full_pleio::copy_sequence_block(), TTQuantiFH::FHread(), TTQFreqExtractor::FHwrite(), TTQOhtaStats::FHwrite(), TTQuanti_continuous_full_pleio::get_additive_genotype(), TTQuanti_diallelic_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_continuous_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_additive_genotype(), TTQuanti_continuous_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), TTQuanti_continuous_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_continuous::init(), TTQuanti_diallelic::init(), TTQuantiSH::init(), TTQuanti_diallelic_bitstring::init(), TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_diallelic_full_pleio::init_sequence(), TTQuanti_diallelic_var_pleio::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio::init_sequence(), TTQuanti_diallelic_bitstring_var_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), TTQuanti_diallelic_full_pleio_epistasis::init_sequence(), TTQuanti_continuous_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::init_sequence(), TTQuanti_continuous::operator==(), TTQuanti_diallelic::operator==(), TTQuanti_diallelic_bitstring::operator==(), TTQuantiFH::print(), TTQuantiFH::print_PLINK_FAM(), TTQuantiFH::print_PLINK_PED(), TTQuanti::reset_phenotype_to_genotypic_value(), LCE_PhenotypeExpression::set_g_value_matrix(), TTQuanti::set_value(), LCE_PhenotypeExpression::setParameters(), LCE_QuantiModifier::setVefromVa(), LCE_Breed_Quanti::setVefromVa(), TTQuanti_continuous_full_pleio::show_up(), TTQuanti_continuous_var_pleio::show_up(), TTQuanti_continuous_no_pleio::show_up(), TTQuanti_diallelic_no_pleio::show_up(), TTQuanti_diallelic_full_pleio::show_up(), TTQuanti_diallelic_var_pleio::show_up(), TTQuanti_diallelic_bitstring_no_pleio::show_up(), TTQuanti_diallelic_bitstring_full_pleio::show_up(), TTQuanti_diallelic_bitstring_var_pleio::show_up(), TTQuanti_continuous_full_pleio_epistasis::show_up(), TTQuanti_diallelic_full_pleio_epistasis::show_up(), TTQuanti_continuous_no_pleio_epistasis::show_up(), TTQuanti_diallelic_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::show_up(), TTQuantiFH::write_PLINK(), and TTQuantiFH::write_TABLE().
◆ get_phenotype_dimension()
|
inlinevirtual |
Returns the dimension of the phenotype of the trait (size of the array accessed with TTrait::getValue()
Implements TraitPrototype.
References _num_traits.
◆ get_phenotypic_value()
|
inline |
References _set_trait_value_func_ptr.
Referenced by TTQuanti::set_value().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_pleio_matrix()
|
inline |
References _pleio_matx.
◆ get_pleiotropy_type()
|
inline |
References _pleio_type.
Referenced by TTQuanti_continuous::operator==(), TTQuanti_diallelic::operator==(), and TTQuanti_diallelic_bitstring::operator==().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_seq_diallele_value()
|
inline |
References _sequence_diallele_values.
Referenced by TTQuantiFH::FHread(), TTQuanti_diallelic_no_pleio::get_additive_genotype(), TTQuanti_diallelic_full_pleio::get_additive_genotype(), TTQuanti_diallelic_var_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_additive_genotype(), TTQuanti_diallelic::get_allele_value(), TTQuanti_diallelic_bitstring::get_allele_value(), TTQuanti_diallelic_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype(), TTQuanti_diallelic_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::get_dominant_genotype(), TTQuanti_diallelic::set_allele_value(), and TTQuanti_diallelic_bitstring::set_allele_value().
◆ get_seq_length()
|
inline |
References _seq_length.
Referenced by TTQuantiFH::FHread(), TTQFreqExtractor::FHwrite(), TTQuanti_diallelic::get_allele(), TTQuanti_diallelic_bitstring::get_allele(), TTQuanti_continuous::get_allele_value(), TTQuanti_diallelic::get_allele_value(), TTQuanti_diallelic_bitstring::get_allele_value(), TTQuantiSH::getSNPalleleFreqInPatch(), TTQuanti_continuous::init(), TTQuanti_diallelic::init(), TTQuanti_diallelic_bitstring::init(), TTQuanti_diallelic_no_pleio::init_sequence(), TTQuanti_diallelic_full_pleio::init_sequence(), TTQuanti_diallelic_var_pleio::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio::init_sequence(), TTQuanti_diallelic_bitstring_var_pleio::init_sequence(), TTQuanti_diallelic_full_pleio_epistasis::init_sequence(), TTQuanti_diallelic_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::init_sequence(), TTQuanti_continuous::operator=(), TTQuanti_diallelic::operator=(), TTQuanti_continuous::operator==(), TTQuanti_diallelic::operator==(), TTQuanti_diallelic_bitstring::operator==(), TTQuanti_continuous::retrieve_data(), TTQuanti_diallelic::retrieve_data(), TTQuanti_diallelic_bitstring::set_allele_bit(), TTQuanti_continuous::set_sequence(), TTQuanti_diallelic::set_sequence(), TTQuanti_continuous_full_pleio::show_up(), TTQuanti_continuous_var_pleio::show_up(), TTQuanti_continuous_no_pleio::show_up(), TTQuanti_diallelic_no_pleio::show_up(), TTQuanti_diallelic_full_pleio::show_up(), TTQuanti_diallelic_var_pleio::show_up(), TTQuanti_diallelic_bitstring_no_pleio::show_up(), TTQuanti_diallelic_bitstring_full_pleio::show_up(), TTQuanti_diallelic_bitstring_var_pleio::show_up(), TTQuanti_continuous_full_pleio_epistasis::show_up(), TTQuanti_diallelic_full_pleio_epistasis::show_up(), TTQuanti_continuous_no_pleio_epistasis::show_up(), TTQuanti_diallelic_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::show_up(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::show_up(), TTQuanti_continuous::store_data(), and TTQuanti_diallelic::store_data().
◆ get_sequence_block_size()
|
inline |
References _locus_table, _num_locus, and _seq_length.
Referenced by TTQuanti_diallelic_bitstring_var_pleio::copy_sequence_block(), TTQuanti_continuous_var_pleio::copy_sequence_block(), TTQuanti_diallelic_var_pleio::copy_sequence_block(), TTQuanti_continuous_var_pleio::get_sequence_block_size(), TTQuanti_diallelic_var_pleio::get_sequence_block_size(), and TTQuanti_diallelic_bitstring_var_pleio::get_sequence_block_size().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_size_locus_type()
|
inline |
References _sizeofLocusType.
Referenced by TTQuanti_continuous_var_pleio::copy_sequence_1locus(), TTQuanti_diallelic_var_pleio::copy_sequence_1locus(), TTQuanti_continuous_var_pleio::copy_sequence_block(), TTQuanti_diallelic_var_pleio::copy_sequence_block(), TTQuanti_continuous_var_pleio::get_sequence_block_size(), and TTQuanti_diallelic_var_pleio::get_sequence_block_size().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_stater()
|
inline |
References _stats.
Referenced by LCE_QuantiModifier::setVefromVa(), and LCE_Breed_Quanti::setVefromVa().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_trait_mask()
|
inline |
References _trait_masks.
Referenced by TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), and TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ get_trait_mutation_variance()
| double TProtoQuanti::get_trait_mutation_variance | ( | unsigned int | trait | ) |
References _genomic_mutation_rate, _mutation_rate, _mutation_sigma, TMatrix::colSum(), TMatrix::get(), and TMatrix::nrows().
◆ get_type()
|
inlinevirtual |
Implements TraitPrototype.
References QUANT.
Referenced by TTQuantiSH::init(), TTQuantiFH::setOutputOption(), and TTQuantiFH::write_PLINK().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectBivariateDiallelic()
| double * TProtoQuanti::getMutationEffectBivariateDiallelic | ( | unsigned int | loc | ) |
References _allele_value, _effects_bivar, _mutation_correlation, TMatrix::get(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectBivariateGaussian()
| double * TProtoQuanti::getMutationEffectBivariateGaussian | ( | unsigned int | loc | ) |
References _effects_bivar, _mutation_correlation, _mutation_sigma, RAND::BivariateGaussian(), and TMatrix::get().
Referenced by setMutationModel_full_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectBivariateGaussianLocSpec()
| double * TProtoQuanti::getMutationEffectBivariateGaussianLocSpec | ( | unsigned int | loc | ) |
References _effects_bivar, _mutation_correlation, _mutation_sigma, RAND::BivariateGaussian(), and TMatrix::get().
Referenced by setMutationModel_full_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectMultivariateGaussian()
| double * TProtoQuanti::getMutationEffectMultivariateGaussian | ( | unsigned int | loc | ) |
References _effects_multivar, _eval, _evect, _ws, and get_locus_PD().
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectMultivariateGaussianLocSpec()
| double * TProtoQuanti::getMutationEffectMultivariateGaussianLocSpec | ( | unsigned int | loc | ) |
References _effects_multivar, _eval, _evect, _ws, and get_locus_PD().
Referenced by setMutationModel_full_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffects()
|
inline |
References _getMutationValues.
Referenced by TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), and TTQuanti_continuous_no_pleio_epistasis::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectsVarPleio()
|
inline |
References _getMutationValuesVarPleio.
Referenced by TTQuanti_continuous_var_pleio::init_sequence().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectUnivariateGaussian()
| double * TProtoQuanti::getMutationEffectUnivariateGaussian | ( | unsigned int | loc | ) |
References _effects_bivar, _mutation_sigma, RAND::Gaussian(), and TMatrix::get().
Referenced by setMutationModel_no_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMutationEffectUnivariateGaussianLocSpec()
| double * TProtoQuanti::getMutationEffectUnivariateGaussianLocSpec | ( | unsigned int | loc | ) |
References _effects_bivar, _mutation_sigma, RAND::Gaussian(), and TMatrix::get().
Referenced by setMutationModel_no_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ has_equal_domCoeff()
|
inline |
References _equal_dom_coeff.
Referenced by TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), and TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ has_equal_effects()
|
inline |
References _equal_effects.
Referenced by TTQuanti_diallelic_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_var_pleio::get_additive_genotype(), TTQuanti_diallelic_bitstring_no_pleio::get_dominant_genotype(), TTQuanti_diallelic_bitstring_full_pleio::get_dominant_genotype(), and TTQuanti_diallelic_bitstring_var_pleio::get_dominant_genotype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ hatch()
|
virtual |
Implements TraitPrototype.
References _allele_model, _diallele_datatype, _epistasis, _pleio_type, fatal(), TTQuanti::set_proto(), TTQuanti_continuous_full_pleio, TTQuanti_continuous_no_pleio, and TTQuanti_continuous_var_pleio.
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ inherit()
References _inherit_fun_ptr.
Referenced by TTQuanti::inherit().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ inherit_free()
References _num_locus, TTQuanti::copy_sequence_1locus(), and RAND::RandBool().
Referenced by setGeneticMapParams().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ inherit_low()
References TTProtoWithMap::_map, TTProtoWithMap::_mapIndex, _sizeofLocusType, TTQuanti::copy_sequence_block(), and GeneticMap::reduceJunctions().
Referenced by setGeneticMapParams().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ loadFileServices()
|
virtual |
Implements SimComponent.
References _allele_model, _freqExtractor, _num_traits, _ohtaStats, _reader, _writer, FileServices::attach(), FileServices::attach_reader(), error(), fatal(), get_allele_model(), SimComponent::get_parameter(), Param::getArg(), Param::getMatrix(), Param::getValue(), Param::isMatrix(), Param::isSet(), SIMenv::MainSim, TTQFreqExtractor::resetTable(), TraitFileHandler< TP >::set(), FileHandler::set_isInputHandler(), TraitFileHandler< TP >::set_multi(), and TTQuantiFH::setOutputOption().
◆ loadStatServices()
|
virtual |
Implements SimComponent.
References _stats, and StatServices::attach().
◆ mutate()
|
inline |
References _mutation_func_ptr.
Referenced by TTQuanti::mutate().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_diallelic_no_pleio()
|
inline |
References _num_locus, get_num_mutations(), TTQuanti::mutate_inplace(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_no_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_diallelic_pleio()
|
inline |
References _mutation_correlation, _num_locus, _num_traits, TMatrix::get(), TTQuanti::get_allele_bit(), get_num_mutations(), TTQuanti::mutate_inplace(), RAND::RandBool(), TTQuanti::set_allele_bit(), and RAND::Uniform().
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_diallelic_var_pleio()
|
inline |
References _locus_table, _mutation_correlation, _num_locus, TMatrix::get(), TTQuanti::get_allele_bit(), get_num_mutations(), TTQuanti::mutate_inplace(), RAND::RandBool(), TTQuanti::set_allele_bit(), and RAND::Uniform().
Referenced by setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_full_pleio()
|
inline |
References _getMutationValues, _num_locus, _num_traits, get_num_mutations(), TTQuanti::mutate_add(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_inplace_full_pleio()
|
inline |
References _getMutationValues, _num_locus, _num_traits, get_num_mutations(), TTQuanti::mutate_inplace(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_inplace_no_pleio()
|
inline |
References _getMutationValues, _num_locus, get_num_mutations(), TTQuanti::mutate_inplace(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_no_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_inplace_var_pleio()
|
inline |
References _getMutationValuesVarPleio, _locus_table, _num_locus, get_num_mutations(), TTQuanti::mutate_inplace(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_nill()
|
inline |
Referenced by setMutationModel_full_pleio(), setMutationModel_no_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_no_pleio()
|
inline |
References _getMutationValues, _num_locus, get_num_mutations(), TTQuanti::mutate_add(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_no_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ mutate_var_pleio()
|
inline |
References _getMutationValuesVarPleio, _locus_table, _num_locus, get_num_mutations(), TTQuanti::mutate_add(), RAND::RandBool(), and RAND::Uniform().
Referenced by setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ readMatrixFromQuantiMutationMatrix()
| bool TProtoQuanti::readMatrixFromQuantiMutationMatrix | ( | vector< vector< double >> & | varmat | ) |
References _mutationVarianceIsLocusSpecific, _num_locus, _num_traits, _pleio_type, error(), SimComponent::get_parameter(), Param::getVariableMatrix(), and warning().
Referenced by setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ reset()
|
inlinevirtual |
◆ resetParameterFromSource()
|
inlinevirtual |
Implements SimComponent.
◆ retrieve_data()
|
inlinevirtual |
Implements StorableComponent.
References _seq_length, and BinaryStorageBuffer::read().
◆ set_eVarianceSD()
| void TProtoQuanti::set_eVarianceSD | ( | unsigned int | trait, |
| double | SD | ||
| ) |
References _eVariance.
Referenced by LCE_QuantiModifier::setVefromVa(), and LCE_Breed_Quanti::setVefromVa().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_genotype_value_additive()
| double TProtoQuanti::set_genotype_value_additive | ( | const TTQuanti * | ind, |
| const unsigned int | trait | ||
| ) |
References TTQuanti::get_additive_genotype().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_genotype_value_dominance()
| double TProtoQuanti::set_genotype_value_dominance | ( | const TTQuanti * | ind, |
| const unsigned int | trait | ||
| ) |
References TTQuanti::get_dominant_genotype().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_gsl_mutation_matrix()
| gsl_matrix * TProtoQuanti::set_gsl_mutation_matrix | ( | unsigned int | pleio_deg, |
| const vector< double > & | varcov | ||
| ) |
Referenced by set_gsl_mutation_matrix_from_sigma(), setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_gsl_mutation_matrix_from_sigma()
| gsl_matrix * TProtoQuanti::set_gsl_mutation_matrix_from_sigma | ( | unsigned int | loc, |
| unsigned int | pleio_deg | ||
| ) |
References _mutation_correlation, _mutation_sigma, _mutationVarianceIsLocusSpecific, TMatrix::get(), message(), and set_gsl_mutation_matrix().
Referenced by setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_init_values()
| void TProtoQuanti::set_init_values | ( | const double * | values, |
| unsigned int | nval | ||
| ) |
References _init_value, _num_traits, and TMatrix::reset().
Referenced by LCE_QuantiInit::execute().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_mutation_matrix_decomposition()
| void TProtoQuanti::set_mutation_matrix_decomposition | ( | unsigned int | loc, |
| unsigned int | pleio_deg | ||
| ) |
References _effects_multivar, _eval, _evect, _gsl_mutation_matrix, _ws, fatal(), and message().
Referenced by setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_trait_value_func_ptr()
| void TProtoQuanti::set_trait_value_func_ptr | ( | bool | withVe | ) |
References _set_trait_value_func_ptr, set_trait_value_noVE(), and set_trait_value_VE().
Referenced by LCE_QuantiModifier::execute(), and LCE_Breed_Quanti::setVefromVa().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_trait_value_noVE()
|
inline |
References _set_genotype_func_ptr.
Referenced by set_trait_value_func_ptr(), set_trait_value_VE(), and setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_trait_value_VE()
|
inline |
References _eVariance, RAND::Gaussian(), and set_trait_value_noVE().
Referenced by set_trait_value_func_ptr(), and setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setAlleleModel()
| unsigned int TProtoQuanti::setAlleleModel | ( | ) |
References error(), SimComponent::get_parameter(), Param::getArg(), and message().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setContinuousMutationModel_full_pleio()
| bool TProtoQuanti::setContinuousMutationModel_full_pleio | ( | ) |
References _effects_multivar, _eval, _evect, _gsl_mutation_matrix, _mutation_correlation, _mutation_sigma, _mutationVarianceIsLocusSpecific, _num_locus, _num_traits, _ws, allocate_gsl_mutation_matrix_space(), error(), TMatrix::get(), SimComponent::get_parameter(), Param::getMatrix(), message(), TMatrix::ncols(), TMatrix::nrows(), readMatrixFromQuantiMutationMatrix(), TMatrix::reset(), TMatrix::set(), set_gsl_mutation_matrix(), set_gsl_mutation_matrix_from_sigma(), set_mutation_matrix_decomposition(), setMutationSigmaFromQuantiMutationVariance(), TMatrix::show_up(), and warning().
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setContinuousMutationModel_no_pleio()
| bool TProtoQuanti::setContinuousMutationModel_no_pleio | ( | ) |
References _mutation_sigma, _mutationVarianceIsLocusSpecific, error(), SimComponent::get_parameter(), Param::isSet(), message(), TMatrix::nrows(), setMutationSigmaFromQuantiMutationVariance_no_pleio(), and warning().
Referenced by setMutationModel_no_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setContinuousMutationModel_var_pleio()
| bool TProtoQuanti::setContinuousMutationModel_var_pleio | ( | ) |
References _effects_multivar, _eval, _evect, _gsl_mutation_matrix, _locus_table, _mutation_correlation, _mutation_sigma, _mutationVarianceIsLocusSpecific, _num_locus, _num_traits, _ws, allocate_gsl_mutation_matrix_space(), error(), TMatrix::get(), SimComponent::get_parameter(), message(), TMatrix::ncols(), TMatrix::nrows(), readMatrixFromQuantiMutationMatrix(), TMatrix::reset(), TMatrix::set(), set_gsl_mutation_matrix(), set_gsl_mutation_matrix_from_sigma(), set_mutation_matrix_decomposition(), setMutationSigmaFromQuantiMutationVariance(), and warning().
Referenced by setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setDiallelicMutationModel()
| bool TProtoQuanti::setDiallelicMutationModel | ( | ) |
References _allele_model, _allele_value, _equal_effects, _equal_val_0, _equal_val_1, _equal_val_diff, _num_locus, _num_traits, _seq_length, _sequence_diallele_values, _trait_masks, _trait_table, error(), TMatrix::get(), get_locus_ID(), get_locus_seq_pos(), get_num_locus(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), TMatrix::ncols(), TMatrix::nrows(), TMatrix::reset(), TMatrix::set(), TMatrix::set_col(), and warning().
Referenced by setMutationModel_full_pleio(), setMutationModel_no_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setDominanceParameters()
| bool TProtoQuanti::setDominanceParameters | ( | ) |
References _dominance_effects, _dominance_model, _equal_dom_coeff, _num_locus, _num_traits, TMatrix::assign(), TMatrix::copy_recycle(), error(), TMatrix::get(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), message(), TMatrix::ncols(), TMatrix::nrows(), and TMatrix::reset().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setEpistasisParameters()
| bool TProtoQuanti::setEpistasisParameters | ( | ) |
References _epistasis, _epistatic_coefs_indices, _epistatic_coefs_matrix, _num_epi_coefs, _num_locus, TMatrix::copy(), TMatrix::copy_recycle(), error(), SimComponent::get_parameter(), Param::getMatrix(), Param::getVariableMatrix(), TMatrix::length(), nChooseK(), nChooseKVec(), TMatrix::ncols(), TMatrix::nrows(), TMatrix::reset(), and TMatrix::set().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setGeneticMapParams()
| bool TProtoQuanti::setGeneticMapParams | ( | ) |
References _all_chooser, _inherit_fun_ptr, _num_locus, TTProtoWithMap::_recombRate, inherit_free(), inherit_low(), TTProtoWithMap::isRecombinationFree(), and TTProtoWithMap::setGeneticMapParameters().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setHeritabilityParams()
| bool TProtoQuanti::setHeritabilityParams | ( | ) |
References _eVariance, _h2, _h2_isBroad, _h2_setTime, _num_traits, error(), TMatrix::get(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), Param::isSet(), message(), TMatrix::ncols(), TMatrix::nrows(), and warning().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setInitialValuesParams()
| bool TProtoQuanti::setInitialValuesParams | ( | ) |
References _allele_model, _doInitMutation, _init_value, _init_variance, _num_traits, error(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), Param::isMatrix(), Param::isSet(), TMatrix::ncols(), TMatrix::nrows(), and TMatrix::reset().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setMutationCorrelation()
| bool TProtoQuanti::setMutationCorrelation | ( | ) |
References _mutation_correlation, _num_locus, _num_traits, TMatrix::assign(), TMatrix::copy_recycle(), error(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), TMatrix::ncols(), TMatrix::nrows(), TMatrix::reset(), and warning().
Referenced by setMutationModel_full_pleio(), and setMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setMutationModel_full_pleio()
| bool TProtoQuanti::setMutationModel_full_pleio | ( | ) |
References _allele_model, _genomic_mutation_rate, _getMutationValues, _locusByteSize, _mutation_func_ptr, _mutationVarianceIsLocusSpecific, _num_locus, _num_traits, _seq_length, _sizeofLocusType, error(), getMutationEffectBivariateDiallelic(), getMutationEffectBivariateGaussian(), getMutationEffectBivariateGaussianLocSpec(), getMutationEffectMultivariateGaussian(), getMutationEffectMultivariateGaussianLocSpec(), mutate_diallelic_pleio(), mutate_full_pleio(), mutate_inplace_full_pleio(), mutate_nill(), setContinuousMutationModel_full_pleio(), setDiallelicMutationModel(), setMutationCorrelation(), and setTraitAndLocusTables_full_pleio().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setMutationModel_no_pleio()
| bool TProtoQuanti::setMutationModel_no_pleio | ( | ) |
References _allele_model, _genomic_mutation_rate, _getMutationValues, _locusByteSize, _mutation_func_ptr, _mutationVarianceIsLocusSpecific, _num_locus, _num_traits, SimComponent::_paramSet, _seq_length, _sizeofLocusType, TMatrix::assign(), error(), TMatrix::get(), SimComponent::get_parameter(), Param::getMatrix(), getMutationEffectUnivariateGaussian(), getMutationEffectUnivariateGaussianLocSpec(), ParamSet::isSet(), message(), mutate_diallelic_no_pleio(), mutate_inplace_no_pleio(), mutate_nill(), mutate_no_pleio(), TMatrix::ncols(), TMatrix::nrows(), TMatrix::reset(), TMatrix::rowSum(), setContinuousMutationModel_no_pleio(), setDiallelicMutationModel(), setTraitAndLocusTables_no_pleio(), and warning().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setMutationModel_var_pleio()
| bool TProtoQuanti::setMutationModel_var_pleio | ( | ) |
References _allele_model, _genomic_mutation_rate, _getMutationValuesVarPleio, _locus_table, _locusByteSize, _mutation_func_ptr, _mutationVarianceIsLocusSpecific, _num_locus, _num_traits, _pleio_matx, _seq_length, _sizeofLocusType, _trait_locus_table, _trait_table, error(), TMatrix::get(), SimComponent::get_parameter(), Param::getMatrix(), getMutationEffectBivariateGaussian(), getMutationEffectBivariateGaussianLocSpec(), getMutationEffectMultivariateGaussianLocSpec(), getMutationEffectUnivariateGaussian(), getMutationEffectUnivariateGaussianLocSpec(), TMatrix::getNbCols(), TMatrix::getNbRows(), message(), mutate_diallelic_var_pleio(), mutate_inplace_var_pleio(), mutate_nill(), mutate_var_pleio(), TMatrix::ncols(), TMatrix::nrows(), setContinuousMutationModel_var_pleio(), setDiallelicMutationModel(), and setMutationCorrelation().
Referenced by setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setMutationSigmaFromQuantiMutationVariance()
| bool TProtoQuanti::setMutationSigmaFromQuantiMutationVariance | ( | ) |
References _mutation_sigma, _num_locus, _num_traits, error(), TMatrix::get(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), message(), TMatrix::ncols(), TMatrix::nrows(), TMatrix::reset(), TMatrix::set_row(), TMatrix::show_up(), and warning().
Referenced by setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setMutationSigmaFromQuantiMutationVariance_no_pleio()
| bool TProtoQuanti::setMutationSigmaFromQuantiMutationVariance_no_pleio | ( | ) |
References _mutation_sigma, _num_locus, _num_traits, _trait_table, TMatrix::copy_recycle(), error(), TMatrix::get(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getMatrix(), message(), TMatrix::ncols(), TMatrix::nrows(), TMatrix::reset(), TMatrix::set(), TMatrix::show_up(), and warning().
Referenced by setContinuousMutationModel_no_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setParameters()
|
virtual |
Implements SimComponent.
References _2L, _allele_model, _diallele_datatype, _dominance_model, _eVariance, _genomic_mutation_rate, _getGenotypeWithDominance, _locus_table, _mutation_rate, _num_locus, _num_traits, _pleio_type, _seq_length, _set_genotype_func_ptr, _set_trait_value_func_ptr, _sizeofLocusType, _trait_locus_table, _trait_table, deallocate_gsl_mutation_matrix_space(), error(), get_genotype_dominance_k(), SimComponent::get_parameter(), SimComponent::get_parameter_value(), Param::getArg(), hatch(), TTrait::init(), TTrait::init_sequence(), message(), set_genotype_value_additive(), set_genotype_value_dominance(), set_trait_value_noVE(), set_trait_value_VE(), setAlleleModel(), setDominanceParameters(), setEpistasisParameters(), setGeneticMapParams(), setHeritabilityParams(), setInitialValuesParams(), setMutationModel_full_pleio(), setMutationModel_no_pleio(), setMutationModel_var_pleio(), TTrait::show_up(), and warning().
◆ setTraitAndLocusTables_full_pleio()
| void TProtoQuanti::setTraitAndLocusTables_full_pleio | ( | ) |
References _locus_table, _num_locus, _num_traits, _trait_locus_table, and _trait_table.
Referenced by setMutationModel_full_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setTraitAndLocusTables_no_pleio()
| void TProtoQuanti::setTraitAndLocusTables_no_pleio | ( | TMatrix & | mat | ) |
References _locus_table, _num_locus, _num_traits, _trait_locus_table, _trait_table, and TMatrix::get().
Referenced by setMutationModel_no_pleio().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ store_data()
|
inlinevirtual |
Implements StorableComponent.
References _seq_length, and BinaryStorageBuffer::store().
Friends And Related Function Documentation
◆ TTQuanti
|
friend |
◆ TTQuanti_continuous
|
friend |
◆ TTQuanti_continuous_full_pleio
|
friend |
Referenced by hatch().
◆ TTQuanti_continuous_no_pleio
|
friend |
Referenced by hatch().
◆ TTQuanti_continuous_single
|
friend |
◆ TTQuanti_continuous_var_pleio
|
friend |
Referenced by hatch().
Member Data Documentation
◆ _2L
|
protected |
Diploid locus size, to save on useless operations during mutation.
Referenced by get_num_mutations(), and setParameters().
◆ _all_chooser
|
private |
Referenced by setGeneticMapParams(), and ~TProtoQuanti().
◆ _allele_model
|
protected |
◆ _allele_value
|
private |
◆ _diallele_datatype
|
protected |
Referenced by hatch(), and setParameters().
◆ _doInitMutation
|
private |
Referenced by get_doInitMutation(), and setInitialValuesParams().
◆ _dominance_effects
|
private |
Referenced by get_dominance(), get_dominance_effects(), and setDominanceParameters().
◆ _dominance_model
|
private |
Referenced by get_dominance_model(), setDominanceParameters(), and setParameters().
◆ _effects_bivar
|
private |
◆ _effects_multivar
|
private |
Referenced by allocate_gsl_mutation_matrix_space(), deallocate_gsl_mutation_matrix_space(), getMutationEffectMultivariateGaussian(), getMutationEffectMultivariateGaussianLocSpec(), set_mutation_matrix_decomposition(), setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
◆ _epistasis
|
private |
Referenced by do_epistasis(), hatch(), and setEpistasisParameters().
◆ _epistatic_coefs_indices
|
private |
Referenced by get_epi_coef_index(), setEpistasisParameters(), and TProtoQuanti().
◆ _epistatic_coefs_matrix
|
private |
Referenced by get_epi_coefs(), setEpistasisParameters(), and TProtoQuanti().
◆ _equal_dom_coeff
|
private |
Referenced by has_equal_domCoeff(), and setDominanceParameters().
◆ _equal_effects
|
private |
Referenced by has_equal_effects(), setDiallelicMutationModel(), and TProtoQuanti().
◆ _equal_val_0
|
private |
Referenced by get_equal_val_0(), setDiallelicMutationModel(), and TProtoQuanti().
◆ _equal_val_1
|
private |
Referenced by get_equal_val_1(), setDiallelicMutationModel(), and TProtoQuanti().
◆ _equal_val_diff
|
private |
Referenced by get_equal_val_diff(), setDiallelicMutationModel(), and TProtoQuanti().
◆ _eval
|
private |
Referenced by allocate_gsl_mutation_matrix_space(), deallocate_gsl_mutation_matrix_space(), getMutationEffectMultivariateGaussian(), getMutationEffectMultivariateGaussianLocSpec(), set_mutation_matrix_decomposition(), setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
◆ _eVariance
|
private |
Referenced by get_env_var(), set_eVarianceSD(), set_trait_value_VE(), setHeritabilityParams(), and setParameters().
◆ _evect
|
private |
Referenced by allocate_gsl_mutation_matrix_space(), deallocate_gsl_mutation_matrix_space(), getMutationEffectMultivariateGaussian(), getMutationEffectMultivariateGaussianLocSpec(), set_mutation_matrix_decomposition(), setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
◆ _freqExtractor
|
private |
Referenced by loadFileServices(), and ~TProtoQuanti().
◆ _genomic_mutation_rate
|
protected |
◆ _getGenotypeWithDominance
|
private |
Pointer to either dominance_h() or dominance_k() function computing the genotypic value with dominance.
Referenced by setParameters().
◆ _getMutationValues
|
private |
Pointer to mutation allele value function, which depends on allele model and number of traits affected.
Referenced by getMutationEffects(), TTQuanti_continuous_full_pleio::init_sequence(), mutate_full_pleio(), mutate_inplace_full_pleio(), mutate_inplace_no_pleio(), mutate_no_pleio(), setMutationModel_full_pleio(), and setMutationModel_no_pleio().
◆ _getMutationValuesVarPleio
|
private |
Collection of pointers to mutation functions, which generate allele values in dependence of pleiotropic degree.
Referenced by getMutationEffectsVarPleio(), mutate_inplace_var_pleio(), mutate_var_pleio(), and setMutationModel_var_pleio().
◆ _gsl_mutation_matrix
|
private |
◆ _h2
|
private |
Referenced by get_heritability(), and setHeritabilityParams().
◆ _h2_isBroad
|
private |
Referenced by get_h2_isBroad(), and setHeritabilityParams().
◆ _h2_setTime
|
private |
Referenced by get_h2_setTime(), and setHeritabilityParams().
◆ _inherit_fun_ptr
Pointer to inheritance functions: either inherit_free() (r=0.5), or inherit_low() (r<0.5).
Referenced by inherit(), and setGeneticMapParams().
◆ _init_value
|
private |
Referenced by get_init_value(), set_init_values(), and setInitialValuesParams().
◆ _init_variance
|
private |
Referenced by get_init_variance(), and setInitialValuesParams().
◆ _locus_table
|
private |
Locus table, num_locus x 2, first column holds the start position of the alleles of each locus in the sequence, second column counts the number of alleles = pleiotropic degree.
Allelic values at a locus are contiguous in the sequence.
Referenced by get_locus_PD(), get_locus_start_pos(), get_sequence_block_size(), mutate_diallelic_var_pleio(), mutate_inplace_var_pleio(), mutate_var_pleio(), setContinuousMutationModel_var_pleio(), setMutationModel_var_pleio(), setParameters(), setTraitAndLocusTables_full_pleio(), and setTraitAndLocusTables_no_pleio().
◆ _locusByteSize
|
private |
◆ _mutation_correlation
|
private |
Referenced by getMutationEffectBivariateDiallelic(), getMutationEffectBivariateGaussian(), getMutationEffectBivariateGaussianLocSpec(), mutate_diallelic_pleio(), mutate_diallelic_var_pleio(), set_gsl_mutation_matrix_from_sigma(), setContinuousMutationModel_full_pleio(), setContinuousMutationModel_var_pleio(), and setMutationCorrelation().
◆ _mutation_func_ptr
|
private |
Pointer to mutation function, which depends on allele on model (HC, noHC, diallelic)
Referenced by mutate(), setMutationModel_full_pleio(), setMutationModel_no_pleio(), and setMutationModel_var_pleio().
◆ _mutation_rate
|
protected |
Referenced by get_num_mutations(), get_trait_mutation_variance(), and setParameters().
◆ _mutation_sigma
|
private |
Referenced by get_trait_mutation_variance(), getMutationEffectBivariateGaussian(), getMutationEffectBivariateGaussianLocSpec(), getMutationEffectUnivariateGaussian(), getMutationEffectUnivariateGaussianLocSpec(), set_gsl_mutation_matrix_from_sigma(), setContinuousMutationModel_full_pleio(), setContinuousMutationModel_no_pleio(), setContinuousMutationModel_var_pleio(), setMutationSigmaFromQuantiMutationVariance(), and setMutationSigmaFromQuantiMutationVariance_no_pleio().
◆ _mutationEffectIsFixedDiAllele
|
private |
◆ _mutationVarianceIsLocusSpecific
|
private |
Referenced by deallocate_gsl_mutation_matrix_space(), readMatrixFromQuantiMutationMatrix(), set_gsl_mutation_matrix_from_sigma(), setContinuousMutationModel_full_pleio(), setContinuousMutationModel_no_pleio(), setContinuousMutationModel_var_pleio(), setMutationModel_full_pleio(), setMutationModel_no_pleio(), and setMutationModel_var_pleio().
◆ _num_epi_coefs
|
private |
Referenced by get_num_epi_coefs(), and setEpistasisParameters().
◆ _num_locus
|
protected |
Total number of loci, for all traits.
Referenced by deallocate_gsl_mutation_matrix_space(), get_locus_number(), get_num_locus(), get_sequence_block_size(), inherit_free(), mutate_diallelic_no_pleio(), mutate_diallelic_pleio(), mutate_diallelic_var_pleio(), mutate_full_pleio(), mutate_inplace_full_pleio(), mutate_inplace_no_pleio(), mutate_inplace_var_pleio(), mutate_no_pleio(), mutate_var_pleio(), readMatrixFromQuantiMutationMatrix(), setContinuousMutationModel_full_pleio(), setContinuousMutationModel_var_pleio(), setDiallelicMutationModel(), setDominanceParameters(), setEpistasisParameters(), setGeneticMapParams(), setMutationCorrelation(), setMutationModel_full_pleio(), setMutationModel_no_pleio(), setMutationModel_var_pleio(), setMutationSigmaFromQuantiMutationVariance(), setMutationSigmaFromQuantiMutationVariance_no_pleio(), setParameters(), setTraitAndLocusTables_full_pleio(), and setTraitAndLocusTables_no_pleio().
◆ _num_traits
|
protected |
Number of traits.
Referenced by get_num_traits(), get_phenotype_dimension(), loadFileServices(), mutate_diallelic_pleio(), mutate_full_pleio(), mutate_inplace_full_pleio(), readMatrixFromQuantiMutationMatrix(), set_init_values(), setContinuousMutationModel_full_pleio(), setContinuousMutationModel_var_pleio(), setDiallelicMutationModel(), setDominanceParameters(), setHeritabilityParams(), setInitialValuesParams(), setMutationCorrelation(), setMutationModel_full_pleio(), setMutationModel_no_pleio(), setMutationModel_var_pleio(), setMutationSigmaFromQuantiMutationVariance(), setMutationSigmaFromQuantiMutationVariance_no_pleio(), setParameters(), setTraitAndLocusTables_full_pleio(), setTraitAndLocusTables_no_pleio(), and TProtoQuanti().
◆ _ohtaStats
|
private |
Referenced by loadFileServices(), and ~TProtoQuanti().
◆ _pleio_matx
|
private |
Pleiotropy matrix provided in input (num locu X num trait).
Gene (row)-trait (column) connectivity matrix.
Referenced by get_pleio_matrix(), setMutationModel_var_pleio(), and TProtoQuanti().
◆ _pleio_type
|
protected |
Referenced by get_pleiotropy_type(), hatch(), readMatrixFromQuantiMutationMatrix(), and setParameters().
◆ _reader
|
private |
Referenced by loadFileServices(), and ~TProtoQuanti().
◆ _seq_length
|
protected |
Total number of allelic values stored in individual sequences, no trait x no locus.
Referenced by get_seq_length(), get_sequence_block_size(), retrieve_data(), setDiallelicMutationModel(), setMutationModel_full_pleio(), setMutationModel_no_pleio(), setMutationModel_var_pleio(), setParameters(), and store_data().
◆ _sequence_diallele_values
|
private |
Referenced by get_seq_diallele_value(), setDiallelicMutationModel(), TProtoQuanti(), and ~TProtoQuanti().
◆ _set_genotype_func_ptr
|
private |
Pointer to functions get_genotype_value_additive() or get_genotype_value_dominance() computing the genotypic value of a trait as function of allele effect.
Referenced by get_genotypic_value(), set_trait_value_noVE(), and setParameters().
◆ _set_trait_value_func_ptr
|
private |
Pointer to either set_trait_value_VE() or set_trait_value_noVE() to compute phenotypic values.
Will call function to compute genotype values stored in _set_genotype_func_ptr.
Referenced by get_phenotypic_value(), set_trait_value_func_ptr(), and setParameters().
◆ _sizeofLocusType
|
private |
◆ _stats
|
private |
Referenced by get_stater(), loadStatServices(), and ~TProtoQuanti().
◆ _trait_locus_table
|
private |
Table storing the locus id of each locus affecting each trait (num trait X (variable length/trait)).
Referenced by get_allele_position(), get_locus_ID(), setMutationModel_var_pleio(), setParameters(), setTraitAndLocusTables_full_pleio(), and setTraitAndLocusTables_no_pleio().
◆ _trait_masks
|
private |
Referenced by get_trait_mask(), and setDiallelicMutationModel().
◆ _trait_table
|
private |
Trait table, (num trait X (variable length/trait)), holds, for each trait, the array position of causative alleles in the sequence.
Referenced by get_allele_position(), get_locus_seq_pos(), get_num_locus(), setDiallelicMutationModel(), setMutationModel_var_pleio(), setMutationSigmaFromQuantiMutationVariance_no_pleio(), setParameters(), setTraitAndLocusTables_full_pleio(), and setTraitAndLocusTables_no_pleio().
◆ _writer
|
private |
Referenced by loadFileServices(), and ~TProtoQuanti().
◆ _ws
|
private |
Referenced by allocate_gsl_mutation_matrix_space(), deallocate_gsl_mutation_matrix_space(), getMutationEffectMultivariateGaussian(), getMutationEffectMultivariateGaussianLocSpec(), set_mutation_matrix_decomposition(), setContinuousMutationModel_full_pleio(), and setContinuousMutationModel_var_pleio().
The documentation for this class was generated from the following files:
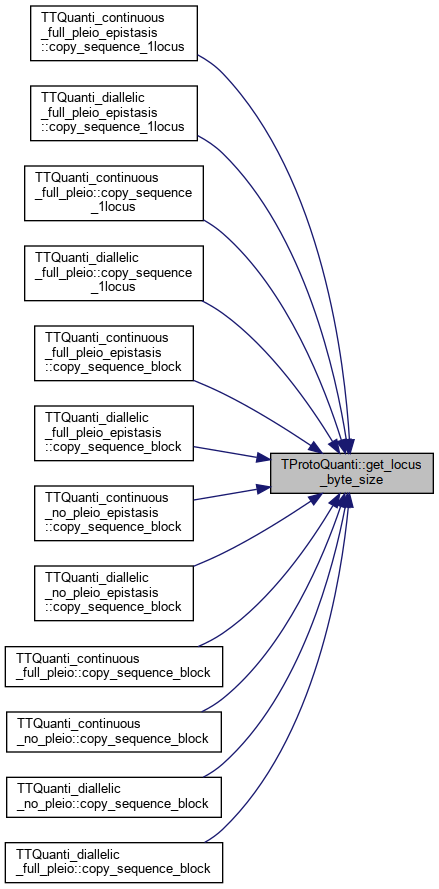
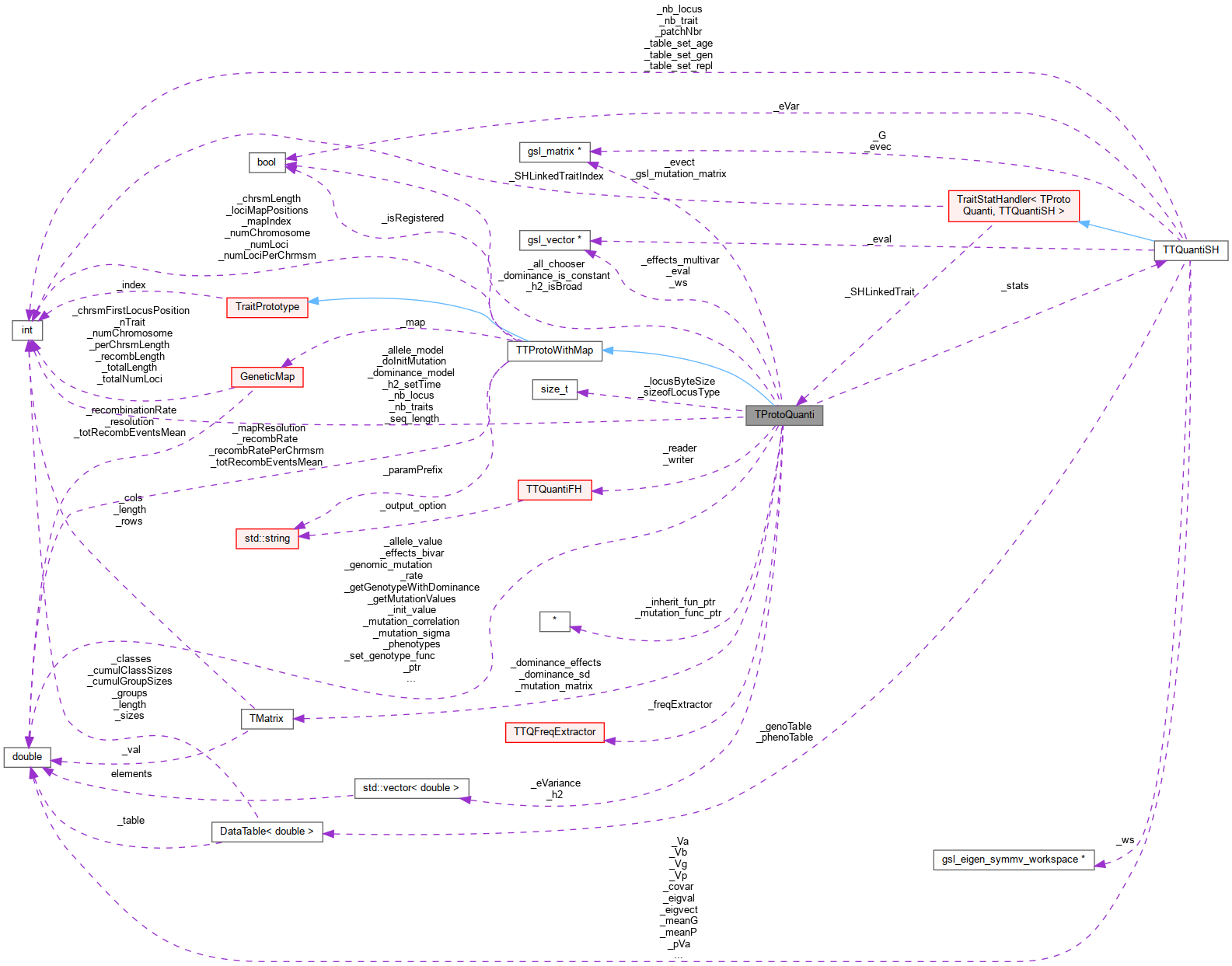






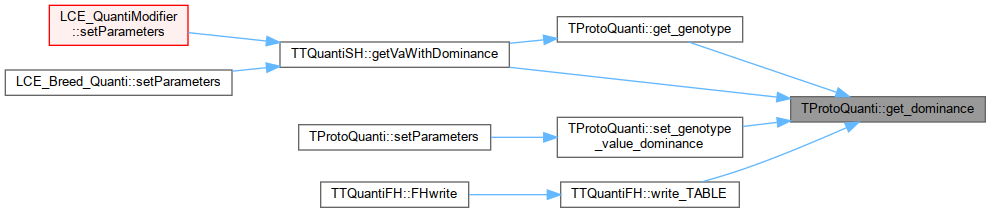

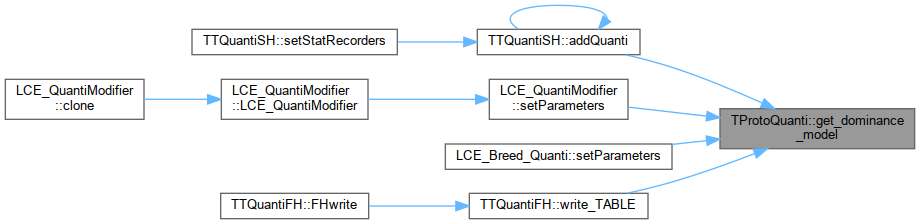


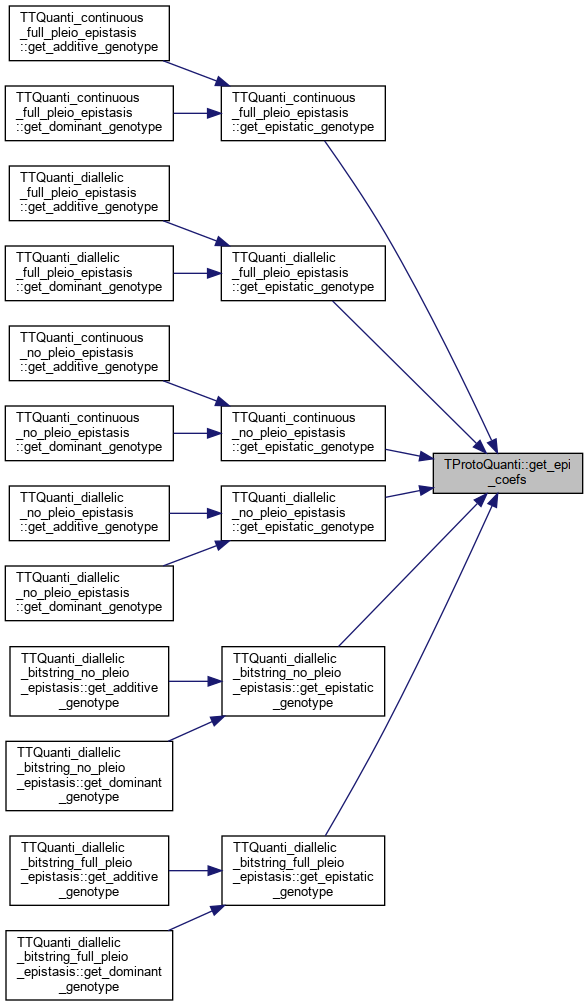

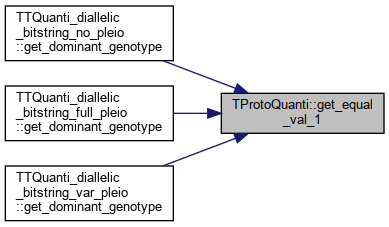
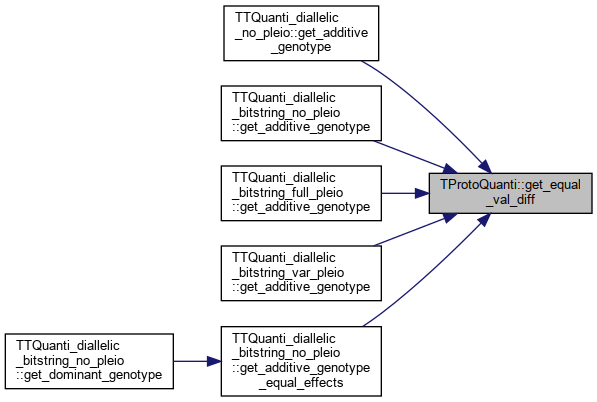

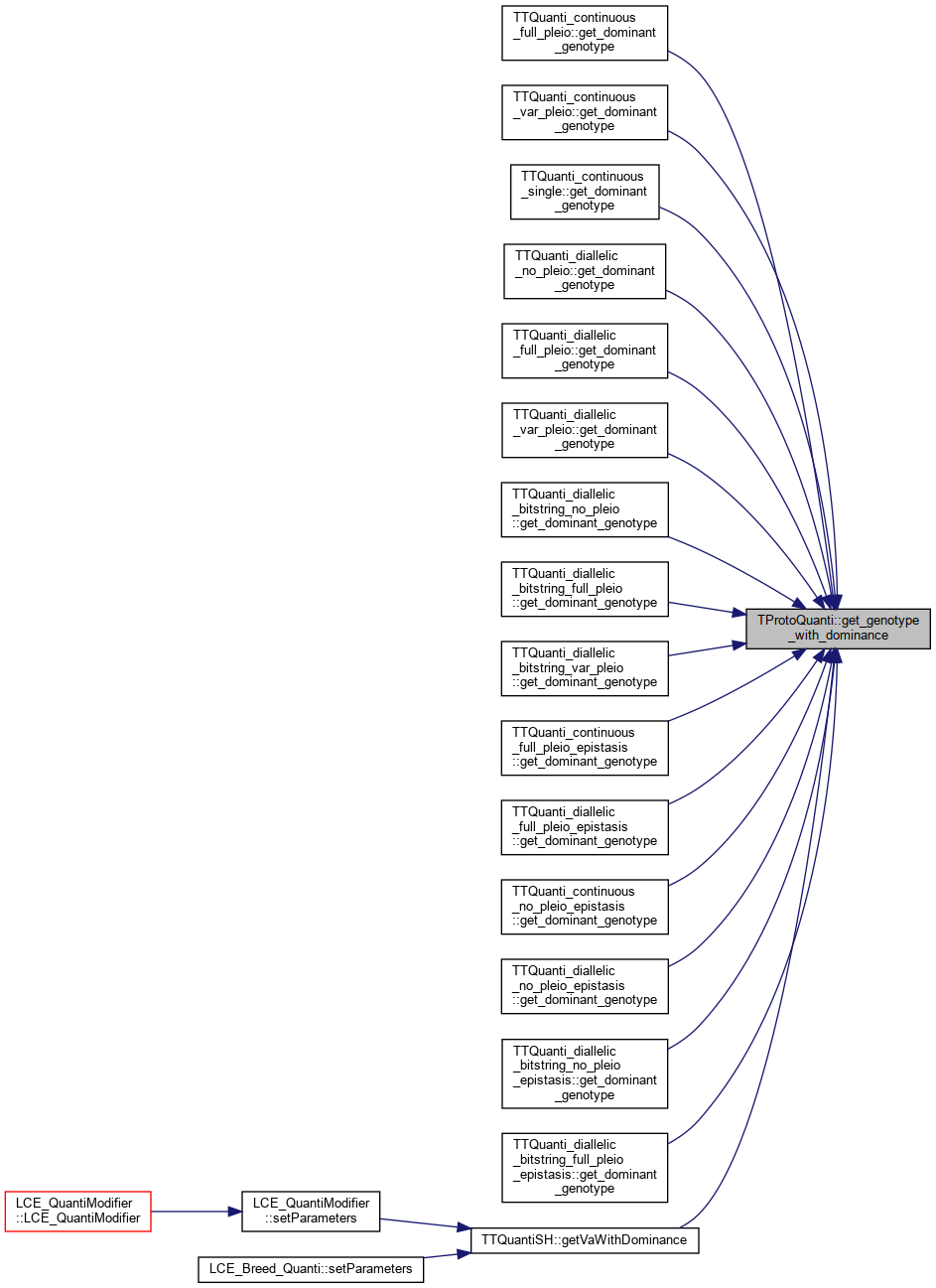
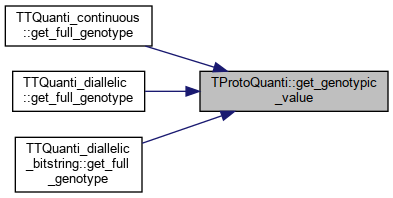


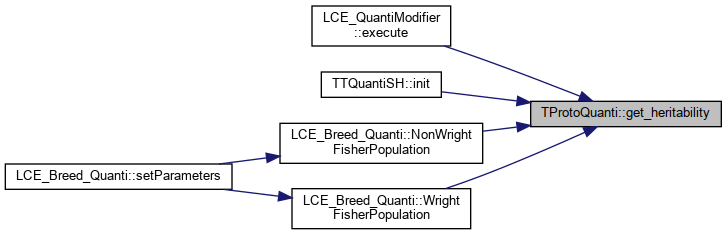


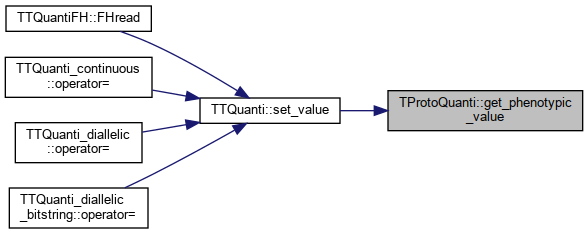
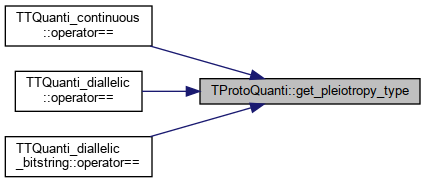

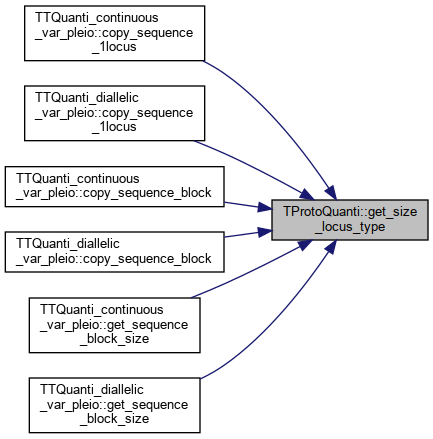
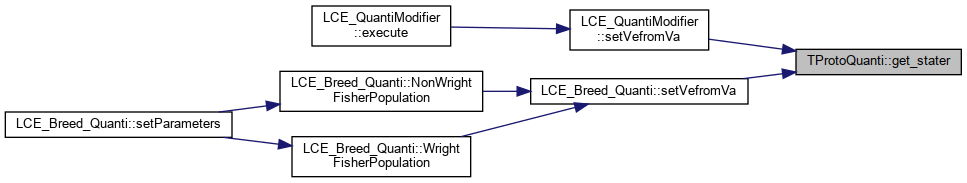
























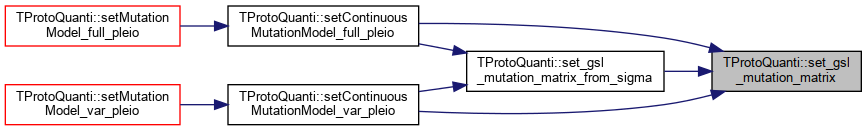
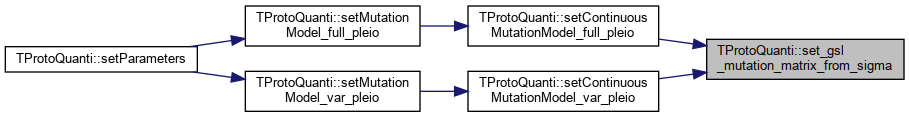
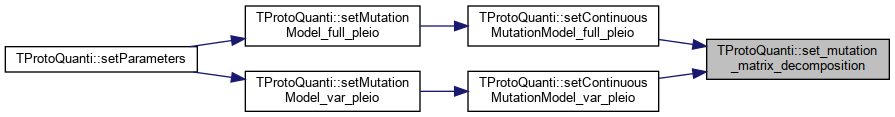

















 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
