Random number generation class, uses various types of random generators depending on the implementation. More...
#include <Uniform.h>
 Collaboration diagram for RAND:
Collaboration diagram for RAND:Static Public Member Functions | |
| static void | init (unsigned long seed) |
| Initialize the random generator's seed. More... | |
| static void | free () |
| Memory de-allocation. More... | |
| static double | Uniform () |
| Generates a random number from [0.0, 1.0[ uniformly distributed. More... | |
| static unsigned int | Uniform (unsigned int max) |
| Returns a uniformly distributed random number from [0.0, max[. More... | |
| static bool | RandBool () |
| Returns a random boolean. More... | |
| static unsigned long | RandULong () |
| Return a random unsigned long, from uniform distribution. More... | |
| static double | gammln (double xx) |
| From the Numerical Recieps. More... | |
| static double | Poisson (double mean) |
| From the Numerical Recieps. More... | |
| static double | Gaussian (double sigma) |
| static void | BivariateGaussian (double sigma1, double sigma2, double rho, double *out1, double *out2) |
| static double | LogNormal (double zeta, double sigma) |
| static double | Gamma (double a, double b) |
| static double | Bernoulli (double p) |
| static double | Exponential (double mu) |
| static double | Binomial (double p, unsigned int n) |
| static double | Beta (const double a, const double b) |
| static unsigned int | Binomial2 (double p, unsigned int n) |
| static void | Multinomial (size_t K, unsigned int N, const double p[], unsigned int n[]) |
| static void | MultinomialOnNormalizedValarray (size_t K, unsigned int N, const std::valarray< double > &p, unsigned int n[]) |
| Multinomial draw assuming the probabilities sum to 1.0 and are all > 0. More... | |
| static void | MultinomialOnNormalizedValarray_expandedOut (size_t K, unsigned int N, const std::valarray< double > &p, unsigned int n[]) |
| Multinomial draw assuming the probabilities sum to 1.0 and are all > 0, the output is an array of size N. More... | |
| static void | MultinomialOnNormalizedValarray_scrambleOut (size_t K, unsigned int N, const std::valarray< double > &p, unsigned int n[]) |
| Multinomial draw assuming the probabilities sum to 1.0 and are all > 0, the output is an array of size N. More... | |
| static void | MultinomialOnNormalizedValarrayZipper_scrambleOut (size_t K, unsigned int N, const std::valarray< double > &p, unsigned int n[]) |
| Multinomial draw assuming the probabilities sum to 1.0 and are all > 0, the output is an array of size N. More... | |
| static void | ScrambleArrayUInt (const int length, unsigned int *array) |
| Randomize the elements within an array. More... | |
| static void | Sample (const int from, const int to, const unsigned int num, int *result, bool replace) |
| Creates a sample of integers within range [from, to), with or without replacement. More... | |
| static void | SampleSeq (int from, int to, int by, unsigned int num, int *result, bool replace=false) |
| static void | SampleSeqWithReciprocal (int from, int to, int by, unsigned int num1, int *result1, unsigned int num2, int *result2) |
| static size_t | Discrete (const gsl_ran_discrete_t *g) |
| Calling the GSL ran_discrete function. More... | |
Static Public Attributes | |
| static long | Seed1 = 0 |
| static long | Seed2 = 98280582 |
Private Member Functions | |
| RAND () | |
Detailed Description
Random number generation class, uses various types of random generators depending on the implementation.
Constructor & Destructor Documentation
◆ RAND()
|
private |
Member Function Documentation
◆ Bernoulli()
|
inlinestatic |
References Uniform().
Referenced by TTNeutralGenes_byte::init_sequence(), and ParamsParser::rbernoul().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Beta()
|
inlinestatic |
◆ Binomial()
|
inlinestatic |
References Beta(), and Uniform().
Referenced by Binomial2(), TProtoNeutralGenes::get_num_mutations(), TProtoQuanti::get_num_mutations(), Multinomial(), MultinomialOnNormalizedValarray(), MultinomialOnNormalizedValarray_expandedOut(), and MultinomialOnNormalizedValarray_scrambleOut().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Binomial2()
|
inlinestatic |
References Binomial().
◆ BivariateGaussian()
|
inlinestatic |
References Uniform().
Referenced by TProtoQuanti::getMutationEffectBivariateGaussian(), and TProtoQuanti::getMutationEffectBivariateGaussianLocSpec().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Discrete()
|
inlinestatic |
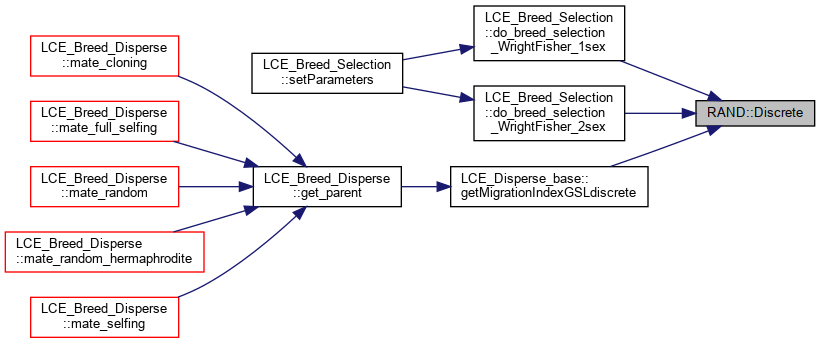
Calling the GSL ran_discrete function.
References Uniform().
Referenced by LCE_Breed_Selection::do_breed_selection_WrightFisher_1sex(), LCE_Breed_Selection::do_breed_selection_WrightFisher_2sex(), and LCE_Disperse_base::getMigrationIndexGSLdiscrete().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Exponential()
|
inlinestatic |
References Uniform().
Referenced by ParamsParser::rexp(), and TProtoDeletMutations_bitstring::set_effects_exp().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ free()
|
inlinestatic |
◆ Gamma()
|
inlinestatic |
References Gaussian(), and Uniform().
Referenced by Beta(), ParamsParser::rgamma(), and TProtoDeletMutations_bitstring::set_effects_gamma().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ gammln()
|
inlinestatic |
◆ Gaussian()
|
inlinestatic |
From the GSL.
References Uniform().
Referenced by Gamma(), LCE_PhenotypeExpression::get_env_cue_noise(), LCE_Selection_base::getFitnessMultivariateGaussian_VE(), LCE_Selection_base::getFitnessUnivariateGaussian_VE(), LCE_Breed_base::getGaussianFecundity(), TProtoQuanti::getMutationEffectUnivariateGaussian(), TProtoQuanti::getMutationEffectUnivariateGaussianLocSpec(), TTDispersal::init_sequence(), TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), TTQuanti_continuous_no_pleio_epistasis::init_sequence(), LCE_Patch_Extinction::rand_gaussian(), ParamsParser::rnorm(), and TProtoQuanti::set_trait_value_VE().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ init()
|
inlinestatic |
Initialize the random generator's seed.
References _myenv, message(), Seed1, MPIenv::workerCount(), and MPIenv::workerRank().
Referenced by SimRunner::init_random_seed(), and SimRunner::run().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ LogNormal()
|
inlinestatic |
References Uniform().
Referenced by LCE_Breed_base::getLogNormalFecundity(), LCE_Patch_Extinction::rand_lognormal(), ParamsParser::rlognorm(), and TProtoDeletMutations_bitstring::set_effects_lognorm().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Multinomial()
|
inlinestatic |
References Binomial().
◆ MultinomialOnNormalizedValarray()
|
inlinestatic |
◆ MultinomialOnNormalizedValarray_expandedOut()
|
inlinestatic |
Multinomial draw assuming the probabilities sum to 1.0 and are all > 0, the output is an array of size N.
References Binomial().
◆ MultinomialOnNormalizedValarray_scrambleOut()
|
inlinestatic |
Multinomial draw assuming the probabilities sum to 1.0 and are all > 0, the output is an array of size N.
References Binomial(), and ScrambleArrayUInt().
◆ MultinomialOnNormalizedValarrayZipper_scrambleOut()
|
inlinestatic |
Multinomial draw assuming the probabilities sum to 1.0 and are all > 0, the output is an array of size N.
References ScrambleArrayUInt(), and Uniform().
◆ Poisson()
|
inlinestatic |
From the Numerical Recieps.
References gammln(), and Uniform().
Referenced by LCE_Breed_base::getPoissonFecundity(), TTDispersal::mutate(), TT_BDMI::mutate_diplo(), TT_BDMI::mutate_haplo(), TTDeletMutations_bitstring::mutate_noredraw(), TTDeletMutations_bitstring::mutate_noredraw_noBackMutation(), TTDeletMutations_bitstring::mutate_redraw(), LCE_Patch_Extinction::rand_poisson(), GeneticMap::recombine(), ParamsParser::rpoiss(), and LCE_Breed_Disperse::stochasticLogisticGrowth().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ RandBool()
|
inlinestatic |
Returns a random boolean.
References RandULong().
Referenced by LCE_Patch_Extinction::do_remove(), TProtoQuanti::getMutationEffectBivariateDiallelic(), LCE_Breed_base::getOffsprgSexFixed(), LCE_Breed_base::getOffsprgSexRandom(), TTDispersal::inherit(), TTNeutralGenes_bitstring::inherit(), TProtoBDMI::inherit_free(), TProtoDeletMutations_bitstring::inherit_free(), TProtoQuanti::inherit_free(), TProtoNeutralGenes::inherit_free(), TTNeutralGenes_bitstring::init_sequence(), TTQuanti_continuous_full_pleio::init_sequence(), TTQuanti_continuous_var_pleio::init_sequence(), TTQuanti_continuous_no_pleio::init_sequence(), TTQuanti_diallelic_no_pleio::init_sequence(), TTQuanti_diallelic_full_pleio::init_sequence(), TTQuanti_diallelic_var_pleio::init_sequence(), TTQuanti_diallelic_bitstring_no_pleio::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio::init_sequence(), TTQuanti_diallelic_bitstring_var_pleio::init_sequence(), TTQuanti_continuous_full_pleio_epistasis::init_sequence(), TTQuanti_diallelic_full_pleio_epistasis::init_sequence(), TTQuanti_continuous_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_no_pleio_epistasis::init_sequence(), TTQuanti_diallelic_bitstring_full_pleio_epistasis::init_sequence(), LCE_Disperse_EvolDisp::Migrate_SteppingStone1D(), TTDispersal::mutate(), TTNeutralGenes_bitstring::mutate(), TTNeutralGenes_byte::mutate_2all(), TProtoQuanti::mutate_diallelic_no_pleio(), TProtoQuanti::mutate_diallelic_pleio(), TProtoQuanti::mutate_diallelic_var_pleio(), TT_BDMI::mutate_diplo(), TProtoQuanti::mutate_full_pleio(), TProtoQuanti::mutate_inplace_full_pleio(), TProtoQuanti::mutate_inplace_no_pleio(), TProtoQuanti::mutate_inplace_var_pleio(), TTNeutralGenes_byte::mutate_KAM(), TProtoQuanti::mutate_no_pleio(), TTDeletMutations_bitstring::mutate_noredraw(), TTDeletMutations_bitstring::mutate_noredraw_noBackMutation(), TTDeletMutations_bitstring::mutate_redraw(), TTNeutralGenes_byte::mutate_SSM(), TProtoQuanti::mutate_var_pleio(), GeneticMap::recombine(), LCE_Breed_Wolbachia::wolbachia_model_1(), and LCE_Breed_Wolbachia::wolbachia_model_2().
◆ RandULong()
|
inlinestatic |
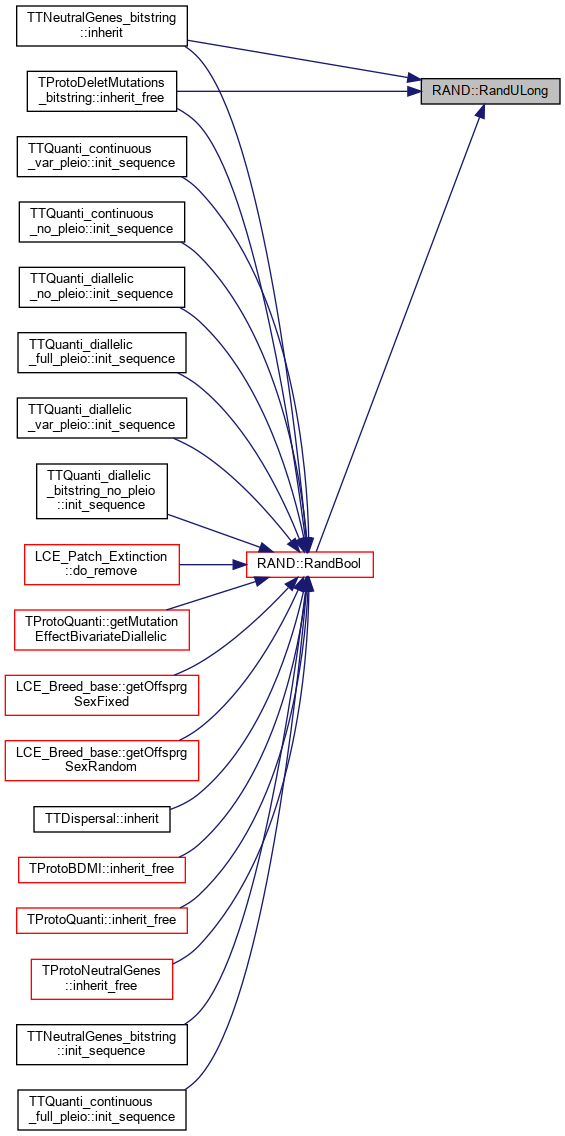
Return a random unsigned long, from uniform distribution.
References Uniform().
Referenced by TTNeutralGenes_bitstring::inherit(), TProtoDeletMutations_bitstring::inherit_free(), and RandBool().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Sample()
|
inlinestatic |
Creates a sample of integers within range [from, to), with or without replacement.
Can be used to scramble an array (without replacement). The random sequence of integers is placed in the 'results' array.
- Parameters
-
from starting number of the series to last number of the series num number of elements to draw within [from, to) result container to hold the resulting randomized sequence of integers
References Uniform().
Referenced by MPFileHandler::createAndPrintSample(), and FileServices::subSamplePatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ SampleSeq()
|
inlinestatic |
References Uniform().
◆ SampleSeqWithReciprocal()
|
inlinestatic |
References Uniform().
Referenced by LCE_NtrlInit::init_allele_freq(), and LCE_QuantiInit::init_allele_freq().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ ScrambleArrayUInt()
|
inlinestatic |
Randomize the elements within an array.
- Parameters
-
length the length of the array array the array to scramble
References Uniform().
Referenced by MultinomialOnNormalizedValarray_scrambleOut(), and MultinomialOnNormalizedValarrayZipper_scrambleOut().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ Uniform() [1/2]
|
inlinestatic |
Generates a random number from [0.0, 1.0[ uniformly distributed.
If SPRNG or GSL libraries are not used, implement a random generator from: L'Ecuyer, 1988, "Efficient and Portable Combined Random Number Generators", Communication of the ACM, 31(6):742-774.
Referenced by Bernoulli(), Binomial(), BivariateGaussian(), LCE_Breed_base::checkPolygyny(), LCE_Cross::create_individual_ancestors(), Discrete(), LCE_Patch_Extinction::do_remove(), LCE_Selection_base::doViabilitySelection(), LCE_Disperse_EvolDisp::evoldisp(), LCE_BreedAssortativeMating::execute(), LCE_Aging::execute(), LCE_Patch_Extinction::execute(), Exponential(), BinaryDataLoader::extractPop(), Metapop::fillPopulationFromSource(), LCE_Disperse_EvolDisp::fixdisp(), LCE_Breed_base::fullPolyginy_manyMales(), Gamma(), Gaussian(), LCE_Breed_Disperse::get_parent(), LCE_Disperse_base::getMigrationIndex(), LCE_Disperse_base::getMigrationPatchBackward(), LCE_Disperse_base::getMigrationPatchForward(), TProtoQuanti::getMutationEffectBivariateDiallelic(), LCE_QuantiInit::init_allele_freq(), TT_BDMI::init_sequence(), TTDeletMutations_bitstring::init_sequence(), TTDispersal::init_sequence(), TTNeutralGenes_byte::init_sequence(), LogNormal(), LCE_Breed_Selection::makeOffspringWithSelection(), LCE_Breed_Disperse::mate_selfing(), LCE_Disperse_EvolDisp::Migrate_Island(), LCE_Disperse_EvolDisp::Migrate_Island_Propagule(), LCE_Disperse_EvolDisp::Migrate_Lattice(), LCE_Disperse_ConstDisp::MigratePatchByNumber(), MultinomialOnNormalizedValarrayZipper_scrambleOut(), TTDispersal::mutate(), TTNeutralGenes_bitstring::mutate(), TTWolbachia::mutate(), TTNeutralGenes_byte::mutate_2all(), TProtoQuanti::mutate_diallelic_no_pleio(), TProtoQuanti::mutate_diallelic_pleio(), TProtoQuanti::mutate_diallelic_var_pleio(), TT_BDMI::mutate_diplo(), TProtoQuanti::mutate_full_pleio(), TT_BDMI::mutate_haplo(), TProtoQuanti::mutate_inplace_full_pleio(), TProtoQuanti::mutate_inplace_no_pleio(), TProtoQuanti::mutate_inplace_var_pleio(), TTNeutralGenes_byte::mutate_KAM(), TProtoQuanti::mutate_no_pleio(), TTDeletMutations_bitstring::mutate_noredraw(), TTDeletMutations_bitstring::mutate_noredraw_noBackMutation(), TTDeletMutations_bitstring::mutate_redraw(), TTNeutralGenes_byte::mutate_SSM(), TProtoQuanti::mutate_var_pleio(), LCE_Breed_base::partialMonoginy(), LCE_Breed_base::partialPolyginy(), LCE_Breed_base::partialPolyginy_manyMales(), LCE_Breed_base::partialSelfing(), Poisson(), LCE_Patch_Extinction::rand_exp(), LCE_Patch_Extinction::rand_uniform(), LCE_Breed_base::random_hermaphrodite(), LCE_Breed_base::RandomMating(), RandULong(), GeneticMap::recombine(), LCE_Resize::regulateAgeClassNoBackup(), LCE_Resize::regulateAgeClassWithBackup(), LCE_Regulation::regulatePatch(), ParamsParser::runif(), Sample(), LCE_Cross::sampleAmongPop(), SampleSeq(), SampleSeqWithReciprocal(), LCE_Cross::sampleWithinPop(), ScrambleArrayUInt(), LCE_BreedAssortativeMating::ScrambleContainer(), LCE_Disperse_base::setPropaguleTargets(), TTProtoWithMap::setRecombinationMapRandom(), setSpatialMatrix(), LCE_Breed_Disperse::stochasticFecundityGrowth(), Uniform(), LCE_Breed_Wolbachia::wolbachia_model_1(), LCE_Breed_base::WrightFisherPopulation(), and LCE_Breed_Quanti::WrightFisherPopulation().
◆ Uniform() [2/2]
|
inlinestatic |
Member Data Documentation
◆ Seed1
◆ Seed2
|
static |
Referenced by Uniform().
The documentation for this class was generated from the following files:















 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
