#include <ttrait_with_map.h>
 Collaboration diagram for GeneticMap:
Collaboration diagram for GeneticMap:Public Member Functions | |
| GeneticMap () | |
| ~GeneticMap () | |
| bool | getGeneticMap (trait_t trait, double **table, unsigned int table_length) |
| double | getResolution () |
| double | setResolution (double val) |
| void | rescaleMap (double val) |
| void | reset_tables () |
| void | clear () |
| void | setLookupTable (unsigned int idx) |
| Bbuilds the lookup table for each trait. More... | |
| void | recombine (sex_t SEX) |
| Called by TTProtoWithMap::recombine twice to create the two gametes necessary for the creation of a new individual. More... | |
| vector< pair< unsigned int, unsigned int > > | reduceJunctions (sex_t SEX, unsigned int trait_idx) |
| Remove multiple x-over at the same locus when traits differ in number of loci. More... | |
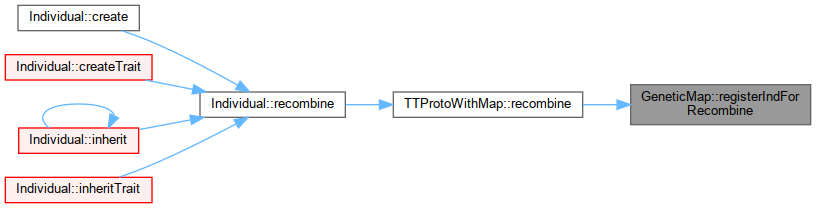
| bool | registerIndForRecombine (unsigned long ID) |
| Called by TTProtoWithMap::recombine with individual ID passed down from Individual::recombine. More... | |
| unsigned int | addTrait (trait_t trait, unsigned int nChrm, unsigned int nLoc, unsigned int *nLocChrm, double resolution, unsigned int *locPositions) |
| Returns the table index for the registered trait. More... | |
| void | unregisterTrait (trait_t trait) |
| bool | checkRegisteredTrait (trait_t trait) |
| Returns true if trait 'trait' has registered a genetic map, false otherwise. More... | |
| vector< unsigned int > & | getRecLoci (sex_t SEX, unsigned int trait) |
| Returns a vector of the loci where crossing-overs take place. More... | |
| vector< bool > & | getFirstRecPosition (sex_t SEX) |
| Returns the vector of the first chromosome position for recombination, used for all traits. More... | |
| unsigned int * | getLocusPositionTable (const trait_t trait) |
Private Attributes | |
| unsigned long | _currentIndividual |
| map< trait_t, unsigned int > | _traits |
| Table mapping trait type to its position index in the following tables. More... | |
| unsigned int | _nTrait |
| Number of traits registered in the map. More... | |
| vector< unsigned int * > | _lociLookupTable |
| A list of tables that map the map position (cM) to a locus, for each trait. More... | |
| vector< unsigned int > | _numChrsmPerTrait |
| Vector of number of chromosomes for each trait. More... | |
| vector< unsigned int > | _numLociPerTrait |
| Vector of number of loci for each trait. More... | |
| vector< unsigned int * > | _numLociPerChrsmPerTrait |
| Vector containing a table of number of loci per chromosome for each trait. More... | |
| vector< unsigned int * > | _locPositionsPerTrait |
| Vector containing the table of map position for the loci of each trait. More... | |
| vector< vector< unsigned int > > | _recPositions [2] |
| Vector of tables containing, for each trait, the locus number at which x-overs happen. More... | |
| vector< bool > | _chrsmFirstRecombPosition [2] |
| Two vectors holding the starting copy of each chromosome to use when creating the two gametes that are used to create a new individual. More... | |
| vector< unsigned int > | _junctions |
| A vector to store the position of the recombination events. More... | |
| unsigned int | _numChromosome |
| unsigned int * | _perChrsmLength |
| unsigned int * | _chrsmFirstLocusPosition |
| unsigned int | _totalLength |
| unsigned int | _recombLength |
| unsigned int | _totalNumLoci |
| double | _resolution |
| double | _totRecombEventsMean |
| double | _recombinationRate |
Detailed Description
Constructor & Destructor Documentation
◆ GeneticMap()
|
inline |
◆ ~GeneticMap()
|
inline |
References reset_tables().
Member Function Documentation
◆ addTrait()
| unsigned int GeneticMap::addTrait | ( | trait_t | trait, |
| unsigned int | nChrm, | ||
| unsigned int | nLoc, | ||
| unsigned int * | nLocChrm, | ||
| double | resolution, | ||
| unsigned int * | locPositions | ||
| ) |
Returns the table index for the registered trait.
References _chrsmFirstLocusPosition, _lociLookupTable, _locPositionsPerTrait, _nTrait, _numChromosome, _numChrsmPerTrait, _numLociPerChrsmPerTrait, _numLociPerTrait, _perChrsmLength, _recombinationRate, _recombLength, _recPositions, _resolution, _totalLength, _totalNumLoci, _totRecombEventsMean, _traits, fatal(), rescaleMap(), reset_tables(), setLookupTable(), and warning().
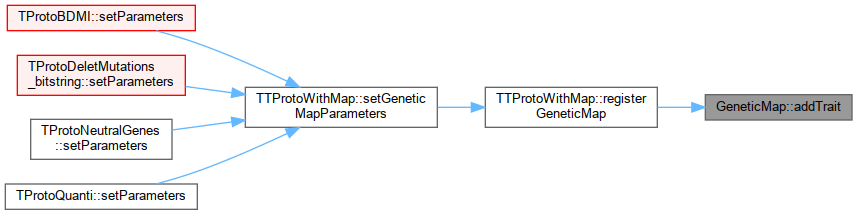
Referenced by TTProtoWithMap::registerGeneticMap().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ checkRegisteredTrait()
| bool GeneticMap::checkRegisteredTrait | ( | trait_t | trait | ) |
Returns true if trait 'trait' has registered a genetic map, false otherwise.
- Parameters
-
trait The trait name to check for in the trait table.
References _traits.
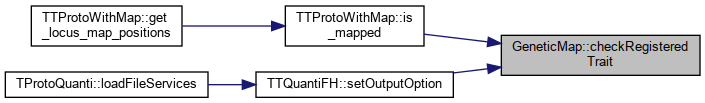
Referenced by TTProtoWithMap::is_mapped(), and TTQuantiFH::setOutputOption().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ clear()
| void GeneticMap::clear | ( | ) |
References _nTrait, _traits, and reset_tables().
Referenced by IndFactory::clearPrototype().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getFirstRecPosition()
|
inline |
Returns the vector of the first chromosome position for recombination, used for all traits.
References _chrsmFirstRecombPosition.
◆ getGeneticMap()
| bool GeneticMap::getGeneticMap | ( | trait_t | trait, |
| double ** | table, | ||
| unsigned int | table_length | ||
| ) |
References _locPositionsPerTrait, _numLociPerChrsmPerTrait, _numLociPerTrait, _traits, and error().
Referenced by TTNeutralGenesFH::write_PLINK(), and TTQuantiFH::write_PLINK().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getLocusPositionTable()
|
inline |
References _locPositionsPerTrait, and _traits.
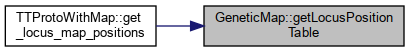
Referenced by TTProtoWithMap::get_locus_map_positions().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getRecLoci()
|
inline |
Returns a vector of the loci where crossing-overs take place.
References _recPositions.
◆ getResolution()
|
inline |
References _resolution.
Referenced by TTProtoWithMap::recordRandomMap().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ recombine()
| void GeneticMap::recombine | ( | sex_t | SEX | ) |
Called by TTProtoWithMap::recombine twice to create the two gametes necessary for the creation of a new individual.
This function sets the location of the x-over on the chromosome map and records the loci positions at which x-over happen for each trait separately. The positions are stored in GeneticMap::_recPositions, which are then passed to the trait-specific inheritance functions when setting up the genetics of the traits in new individuals.
- Parameters
-
SEX the origin of the gamete, whether from the father or the mother
References _chrsmFirstRecombPosition, _junctions, _lociLookupTable, _nTrait, _numChromosome, _numLociPerTrait, _recombLength, _recPositions, _totalNumLoci, _totRecombEventsMean, RAND::Poisson(), RAND::RandBool(), and RAND::Uniform().
Referenced by TTProtoWithMap::recombine().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ reduceJunctions()
| vector< pair< unsigned int, unsigned int > > GeneticMap::reduceJunctions | ( | sex_t | SEX, |
| unsigned int | trait_idx | ||
| ) |
Remove multiple x-over at the same locus when traits differ in number of loci.
References _chrsmFirstRecombPosition, _numChromosome, _numLociPerChrsmPerTrait, _recPositions, and error().
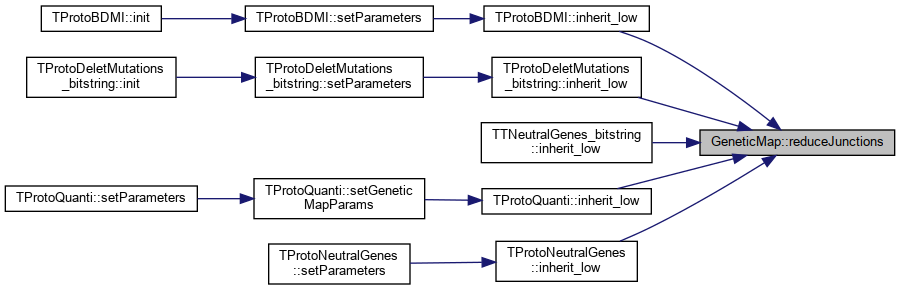
Referenced by TProtoBDMI::inherit_low(), TProtoDeletMutations_bitstring::inherit_low(), TTNeutralGenes_bitstring::inherit_low(), TProtoQuanti::inherit_low(), and TProtoNeutralGenes::inherit_low().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ registerIndForRecombine()
|
inline |
Called by TTProtoWithMap::recombine with individual ID passed down from Individual::recombine.
Returns false when already called by the same individual. The function is called for each trait separately but recombination must be computed once per individual only. It is so because some functions create new individual's trait separately, for instance in breed_selection when traits under viability selection are created before neutral traits.
- Parameters
-
ID the ID number of the individual calling the recombination function
References _currentIndividual.
Referenced by TTProtoWithMap::recombine().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ rescaleMap()
| void GeneticMap::rescaleMap | ( | double | val | ) |
References _chrsmFirstLocusPosition, _locPositionsPerTrait, _nTrait, _numChromosome, _numLociPerTrait, _perChrsmLength, _resolution, _totalLength, _totRecombEventsMean, and setLookupTable().
Referenced by addTrait().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ reset_tables()
| void GeneticMap::reset_tables | ( | ) |
References _chrsmFirstLocusPosition, _chrsmFirstRecombPosition, _lociLookupTable, _locPositionsPerTrait, _numChrsmPerTrait, _numLociPerChrsmPerTrait, _numLociPerTrait, _perChrsmLength, _recPositions, and error().
Referenced by addTrait(), clear(), and ~GeneticMap().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setLookupTable()
| void GeneticMap::setLookupTable | ( | unsigned int | idx | ) |
Bbuilds the lookup table for each trait.
A lookup table maps a chromosomal position to a locus so that when a x-over is placed at a given map position, the corresponding locus can be directly found. The size of the lookup table depends on the _totalLength and _resolution of the map. The lookup table for a 1M map at the 0.01 cM scale will have 10,000 elements.
- Parameters
-
idx the index of the trait in the table (stored in the trait prototype)
!!! +1 needed here because map must start with 0 !!!!
References _chrsmFirstLocusPosition, _lociLookupTable, _locPositionsPerTrait, _numChromosome, _numLociPerChrsmPerTrait, _numLociPerTrait, _perChrsmLength, _resolution, _totalLength, _totRecombEventsMean, and fatal().
Referenced by addTrait(), and rescaleMap().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setResolution()
|
inline |
References _resolution.
◆ unregisterTrait()
| void GeneticMap::unregisterTrait | ( | trait_t | trait | ) |
References _nTrait, _traits, and fatal().
Referenced by TTProtoWithMap::reset(), and TTProtoWithMap::unregisterFromGeneticMap().
 Here is the caller graph for this function:
Here is the caller graph for this function:Member Data Documentation
◆ _chrsmFirstLocusPosition
|
private |
Referenced by addTrait(), rescaleMap(), reset_tables(), and setLookupTable().
◆ _chrsmFirstRecombPosition
|
private |
Two vectors holding the starting copy of each chromosome to use when creating the two gametes that are used to create a new individual.
Referenced by getFirstRecPosition(), recombine(), reduceJunctions(), and reset_tables().
◆ _currentIndividual
|
private |
Referenced by registerIndForRecombine().
◆ _junctions
|
private |
A vector to store the position of the recombination events.
Referenced by recombine().
◆ _lociLookupTable
|
private |
A list of tables that map the map position (cM) to a locus, for each trait.
The length of the table is the length of the genetic map, which depends on the map resolution (cM by default).
Referenced by addTrait(), recombine(), reset_tables(), and setLookupTable().
◆ _locPositionsPerTrait
|
private |
Vector containing the table of map position for the loci of each trait.
Positions are recorded according to the minimum map resolution as used in the lookup table.
Referenced by addTrait(), getGeneticMap(), getLocusPositionTable(), rescaleMap(), reset_tables(), and setLookupTable().
◆ _nTrait
|
private |
Number of traits registered in the map.
Length of following arrays.
Referenced by addTrait(), clear(), recombine(), rescaleMap(), and unregisterTrait().
◆ _numChromosome
|
private |
Referenced by addTrait(), recombine(), reduceJunctions(), rescaleMap(), and setLookupTable().
◆ _numChrsmPerTrait
|
private |
Vector of number of chromosomes for each trait.
Referenced by addTrait(), and reset_tables().
◆ _numLociPerChrsmPerTrait
|
private |
Vector containing a table of number of loci per chromosome for each trait.
Referenced by addTrait(), getGeneticMap(), reduceJunctions(), reset_tables(), and setLookupTable().
◆ _numLociPerTrait
|
private |
Vector of number of loci for each trait.
Referenced by addTrait(), getGeneticMap(), recombine(), rescaleMap(), reset_tables(), and setLookupTable().
◆ _perChrsmLength
|
private |
Referenced by addTrait(), rescaleMap(), reset_tables(), and setLookupTable().
◆ _recombinationRate
|
private |
Referenced by addTrait().
◆ _recombLength
|
private |
Referenced by addTrait(), and recombine().
◆ _recPositions
|
private |
Vector of tables containing, for each trait, the locus number at which x-overs happen.
Updated at each generation. There is matrix per sex/gamete with one trait per row and a variable number of elements in each row.
Referenced by addTrait(), getRecLoci(), recombine(), reduceJunctions(), and reset_tables().
◆ _resolution
|
private |
Referenced by addTrait(), getResolution(), rescaleMap(), setLookupTable(), and setResolution().
◆ _totalLength
|
private |
Referenced by addTrait(), rescaleMap(), and setLookupTable().
◆ _totalNumLoci
|
private |
Referenced by addTrait(), and recombine().
◆ _totRecombEventsMean
|
private |
Referenced by addTrait(), recombine(), rescaleMap(), and setLookupTable().
◆ _traits
|
private |
Table mapping trait type to its position index in the following tables.
Referenced by addTrait(), checkRegisteredTrait(), clear(), getGeneticMap(), getLocusPositionTable(), and unregisterTrait().
The documentation for this class was generated from the following files:









 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
