A file handler to save the neutral markers genotypes in the FSTAT format (extended). More...
#include <ttneutralgenes.h>
 Inheritance diagram for TTNeutralGenesFH:
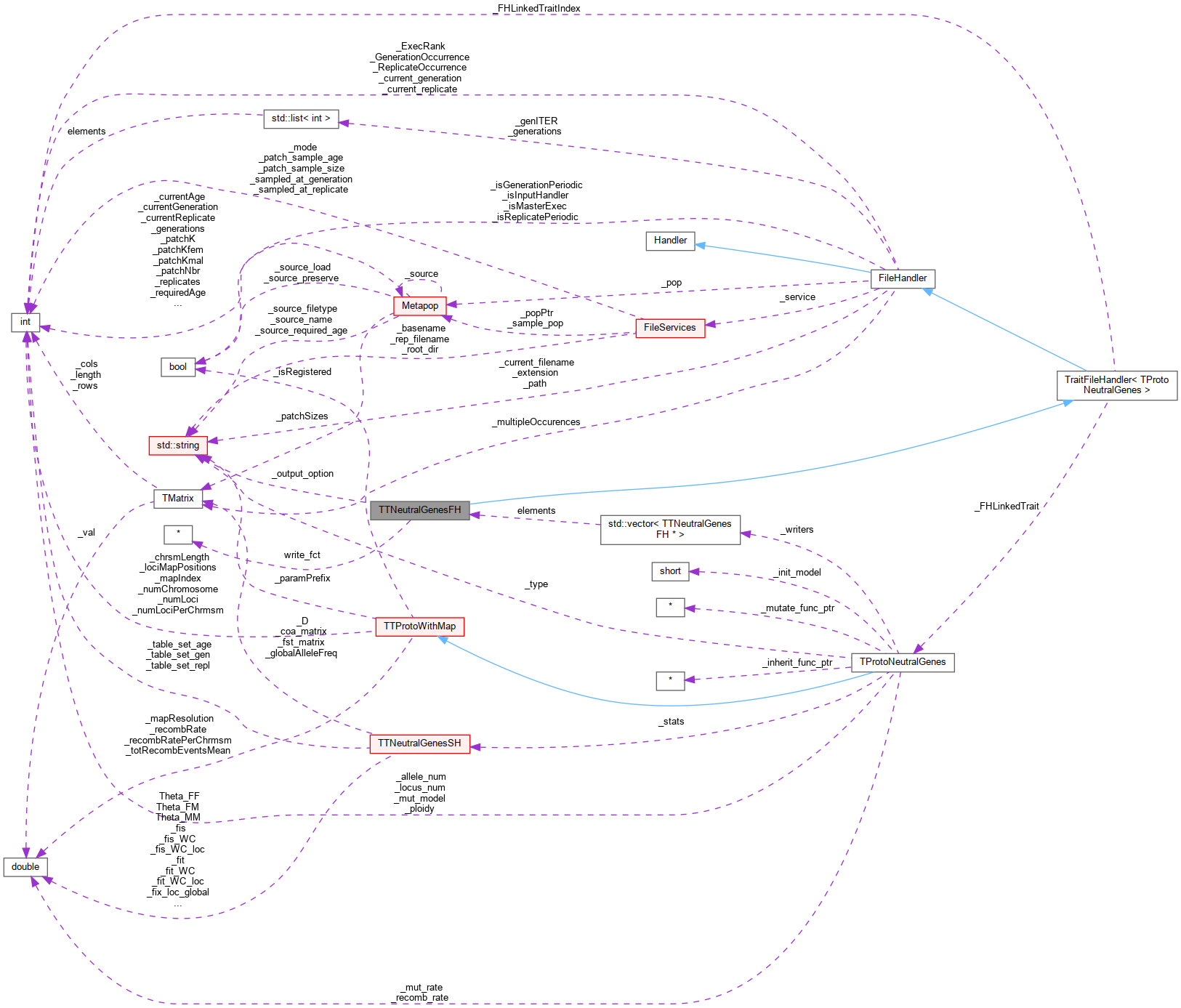
Inheritance diagram for TTNeutralGenesFH: Collaboration diagram for TTNeutralGenesFH:
Collaboration diagram for TTNeutralGenesFH:Public Member Functions | |
| TTNeutralGenesFH (TProtoNeutralGenes *TP) | |
| virtual | ~TTNeutralGenesFH () |
| virtual void | FHwrite () |
| virtual void | FHread (string &filename) |
| void | write_TAB () |
| void | write_patch_TAB (Patch *patch, sex_t SEX, age_idx AGE, ofstream &FH) |
| void | write_PLINK () |
| void | print_PLINK_PED (ofstream &FH, age_idx Ax, Patch *patch) |
| void | write_PLINK_BED (ofstream &BED) |
| void | write_GENEPOP () |
| void | write_patch_GENEPOP (Patch *patch, sex_t SEX, age_idx AGE, ofstream &FH, unsigned int digits) |
| void | write_FSTAT () |
| void | write_patch_FSTAT (Patch *patch, sex_t SEX, age_idx AGE, ofstream &FH, unsigned int digits) |
| void | write_Fst_i () |
| void | write_varcompWC () |
| void | setOutputOption (string opt) |
| void | set_write_fct (void(TTNeutralGenesFH::*fct_ptr)()) |
 Public Member Functions inherited from TraitFileHandler< TProtoNeutralGenes > Public Member Functions inherited from TraitFileHandler< TProtoNeutralGenes > | |
| TraitFileHandler (TProtoNeutralGenes *trait_proto, const char *ext) | |
| virtual | ~TraitFileHandler () |
| virtual void | FHread (string &filename)=0 |
| virtual void | set (bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path, TProtoNeutralGenes *trait_proto) |
| virtual void | set_multi (bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path, TProtoNeutralGenes *trait_proto) |
 Public Member Functions inherited from FileHandler Public Member Functions inherited from FileHandler | |
| FileHandler (const char *ext) | |
| virtual | ~FileHandler () |
| virtual void | init () |
| Called by notifier during simulation setup, performs file checking. More... | |
| virtual vector< string > | ifExist () |
| Checks if any file associated with the current file name already exists on disk. More... | |
| virtual void | set (bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path) |
| Sets the hanlder parameters. More... | |
| virtual void | set_multi (bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path) |
| virtual void | update () |
| Updates the inner replicate and generation counters and calls FHwrite if needed by the the periodicity of the file. More... | |
| Metapop * | get_pop_ptr () |
| Returns the pointer to the current metapop through the FileServices interface. More... | |
| void | set_pop_ptr (Metapop *pop_ptr) |
| FileServices * | get_service () |
| Returns pointer to the FileServices. More... | |
| void | set_service (FileServices *srv) |
| std::string & | get_path () |
| void | set_path () |
| std::string & | get_extension () |
| void | set_extension (const char *ext) |
| std::string & | get_filename () |
| Builds and returns the current file name depending on the periodicity of the file. More... | |
| bool | get_isInputHandler () |
| void | set_isInputHandler (bool val) |
| bool | get_isReplicatePeriodic () |
| void | set_isReplicatePeriodic (bool val) |
| unsigned int | get_ReplicateOccurrence () |
| void | set_ReplicateOccurrence (unsigned int val) |
| bool | get_isGenerationPeriodic () |
| void | set_isGenerationPeriodic (bool val) |
| unsigned int | get_GenerationOccurrence () |
| void | set_GenerationOccurrence (unsigned int val) |
| unsigned int | get_ExecRank () |
| unused yet... More... | |
| void | set_ExecRank (int val) |
| TMatrix * | get_OccMatrix () |
| void | set_OccMatrix (TMatrix *occ) |
| bool | get_isMasterExec () |
| void | set_isMasterExec (bool is) |
 Public Member Functions inherited from Handler Public Member Functions inherited from Handler | |
| virtual | ~Handler () |
Private Attributes | |
| string | _output_option |
| void(TTNeutralGenesFH::* | write_fct )() |
Additional Inherited Members | |
 Protected Attributes inherited from TraitFileHandler< TProtoNeutralGenes > Protected Attributes inherited from TraitFileHandler< TProtoNeutralGenes > | |
| TProtoNeutralGenes * | _FHLinkedTrait |
| int | _FHLinkedTraitIndex |
 Protected Attributes inherited from FileHandler Protected Attributes inherited from FileHandler | |
| Metapop * | _pop |
| Pointer to the current metapop, set during initialization within the init function. More... | |
Detailed Description
A file handler to save the neutral markers genotypes in the FSTAT format (extended).
By default, the file extension is ".dat" for the genotype file. It is changed to ".fsti" for the per-locus/per-patch Weir& Hill Fst's and ".freq" for the per-locus/per-patch allele frequencies. Also implements the FileHandler::FHread method to load a population's genotypes from an FSTAT file (using the 'source_pop' population parameter).
Constructor & Destructor Documentation
◆ TTNeutralGenesFH()
|
inline |
◆ ~TTNeutralGenesFH()
Member Function Documentation
◆ FHread()
|
virtual |
reads in only FSTAT format
Implements FileHandler.
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTraitIndex, FileHandler::_pop, Patch::add(), ADLTx, error(), fatal(), FEM, FSTAT_EXTRA_INFO_LOCI, TProtoNeutralGenes::get_allele_num(), TProtoNeutralGenes::get_locus_num(), Metapop::getPatch(), Metapop::getPatchNbr(), Individual::getTrait(), IndFactory::makeNewIndividual(), message(), OFFSx, TTrait::set_allele_value(), Individual::setPedigreeClass(), and Metapop::size().
◆ FHwrite()
|
virtual |
Implements TraitFileHandler< TProtoNeutralGenes >.
References FileHandler::_pop, FileServices::get_pop_ptr(), FileHandler::get_service(), FileServices::getSampledPop(), Metapop::isAlive(), and write_fct.
◆ print_PLINK_PED()
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTraitIndex, FEM, Patch::get(), TTrait::get_allele(), TProtoNeutralGenes::get_locus_num(), TProtoNeutralGenes::get_ploidy(), Individual::getFatherID(), Individual::getHome(), Individual::getID(), Individual::getMotherID(), Individual::getTrait(), MAL, OFFSx, and Patch::size().
Referenced by write_PLINK().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ set_write_fct()
|
inline |
References write_fct.
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setOutputOption()
| void TTNeutralGenesFH::setOutputOption | ( | string | opt | ) |
References _output_option.
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_Fst_i()
| void TTNeutralGenesFH::write_Fst_i | ( | ) |
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, ADULTS, TTNeutralGenesSH::allocateTables(), fatal(), TProtoNeutralGenes::get_allele_num(), FileHandler::get_extension(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_path(), FileServices::get_pop_ptr(), FileHandler::get_service(), TProtoNeutralGenes::get_stater(), FileServices::getGenerationReplicateFileName(), Metapop::getPatchNbr(), message(), TTNeutralGenesSH::setFst_li(), Metapop::size(), and warning().
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_FSTAT()
| void TTNeutralGenesFH::write_FSTAT | ( | ) |
The file format is FSTAT-like, with extra info about the individual added (age, sex, pedigree, origin). The file extension is ".dat".
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, FileHandler::_pop, ADLTx, fatal(), FEM, FSTAT_EXTRA_INFO_LOCI, TProtoNeutralGenes::get_allele_num(), FileHandler::get_extension(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_path(), TProtoNeutralGenes::get_ploidy(), FileHandler::get_service(), FileServices::getGenerationReplicateFileName(), Metapop::getPatch(), Metapop::getPatchNbr(), MAL, message(), OFFSx, and write_patch_FSTAT().
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_GENEPOP()
| void TTNeutralGenesFH::write_GENEPOP | ( | ) |
The file format is GENEPOP-like, with extra info about the individual added (age, sex, pedigree, origin). The file extension is ".txt".
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, FileHandler::_pop, ADLTx, fatal(), FEM, TProtoNeutralGenes::get_allele_num(), FileHandler::get_extension(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_path(), TProtoNeutralGenes::get_ploidy(), FileHandler::get_service(), FileServices::getGenerationReplicateFileName(), Metapop::getPatch(), Metapop::getPatchNbr(), MAL, message(), OFFSx, and write_patch_GENEPOP().
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_patch_FSTAT()
| void TTNeutralGenesFH::write_patch_FSTAT | ( | Patch * | patch, |
| sex_t | SEX, | ||
| age_idx | AGE, | ||
| ofstream & | FH, | ||
| unsigned int | digits | ||
| ) |
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTraitIndex, Patch::get(), TTrait::get_allele(), TProtoNeutralGenes::get_locus_num(), TProtoNeutralGenes::get_ploidy(), Individual::getHome(), Patch::getID(), Individual::getPedigreeClass(), Individual::getTrait(), and Patch::size().
Referenced by write_FSTAT().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_patch_GENEPOP()
| void TTNeutralGenesFH::write_patch_GENEPOP | ( | Patch * | patch, |
| sex_t | SEX, | ||
| age_idx | AGE, | ||
| ofstream & | FH, | ||
| unsigned int | digits | ||
| ) |
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTraitIndex, Patch::get(), TTrait::get_allele(), TProtoNeutralGenes::get_locus_num(), TProtoNeutralGenes::get_ploidy(), Individual::getHome(), Patch::getID(), Individual::getPedigreeClass(), Individual::getTrait(), and Patch::size().
Referenced by write_GENEPOP().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_patch_TAB()
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTraitIndex, _output_option, Patch::get(), TTrait::get_allele(), TProtoNeutralGenes::get_locus_num(), TProtoNeutralGenes::get_ploidy(), Individual::getFather(), Individual::getFatherID(), Individual::getHome(), Individual::getID(), Patch::getID(), Individual::getMother(), Individual::getMotherID(), Individual::getPedigreeClass(), Individual::getTrait(), and Patch::size().
Referenced by write_TAB().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_PLINK()
| void TTNeutralGenesFH::write_PLINK | ( | ) |
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, TTProtoWithMap::_map, FileHandler::_pop, ADLTx, ADULTS, fatal(), TProtoNeutralGenes::get_allele_num(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_path(), TProtoNeutralGenes::get_ploidy(), FileHandler::get_service(), TProtoNeutralGenes::get_type(), Metapop::getCurrentAge(), FileServices::getGenerationReplicateFileName(), GeneticMap::getGeneticMap(), Metapop::getPatch(), Metapop::getPatchNbr(), message(), OFFSPRG, OFFSx, print_PLINK_PED(), and warning().
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_PLINK_BED()
| void TTNeutralGenesFH::write_PLINK_BED | ( | ofstream & | BED | ) |
◆ write_TAB()
| void TTNeutralGenesFH::write_TAB | ( | ) |
The file extension is ".txt".
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, _output_option, FileHandler::_pop, ADLTx, fatal(), FEM, FileHandler::get_extension(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_path(), TProtoNeutralGenes::get_ploidy(), FileHandler::get_service(), FileServices::getGenerationReplicateFileName(), Metapop::getPatch(), Metapop::getPatchNbr(), MAL, message(), OFFSx, and write_patch_TAB().
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_varcompWC()
| void TTNeutralGenesFH::write_varcompWC | ( | ) |
References TraitFileHandler< TProtoNeutralGenes >::_FHLinkedTrait, _output_option, ADLTx, ALL, TTNeutralGenesSH::allocateTables(), fatal(), DataTable< T >::get(), TMatrix::get(), TProtoNeutralGenes::get_allele_num(), FileHandler::get_extension(), TProtoNeutralGenes::get_locus_num(), FileHandler::get_path(), FileServices::get_pop_ptr(), FileHandler::get_service(), TProtoNeutralGenes::get_stater(), TTNeutralGenesSH::getAlleleCountTable(), TTNeutralGenesSH::getAlleleFreqTable(), Metapop::getCurrentAge(), FileServices::getGenerationReplicateFileName(), TTNeutralGenesSH::getGlobalFreqs(), TTNeutralGenesSH::getHeteroTable(), Metapop::getPatchNbr(), message(), OFFSPRG, OFFSx, TTNeutralGenesSH::setAlleleTables(), TTNeutralGenesSH::setHeteroTable(), TTNeutralGenesSH::setHo2(), and Metapop::size().
Referenced by TProtoNeutralGenes::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:Member Data Documentation
◆ _output_option
|
private |
Referenced by setOutputOption(), write_patch_TAB(), write_TAB(), and write_varcompWC().
◆ write_fct
|
private |
Referenced by FHwrite(), and set_write_fct().
The documentation for this class was generated from the following files:












 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
