|
| | TTNeutralGenesSH (TProtoNeutralGenes *TP) |
| |
| virtual | ~TTNeutralGenesSH () |
| |
| virtual void | init () |
| |
| virtual bool | setStatRecorders (std::string &token) |
| |
| void | setFreqRecorders (age_t AGE) |
| |
| void | setFreqRecordersPerPatch (age_t AGE) |
| |
| void | setFstatRecorders (age_t AGE) |
| |
| void | setFstatWCRecorders (age_t AGE) |
| |
| void | setCoaMatrixRecorders (age_t AGE, unsigned char dim) |
| |
| void | setFstMatrixRecorders (age_t AGE, unsigned char dim) |
| |
| void | setNeiGeneticDistanceRecorders (age_t AGE, bool pairwise) |
| |
| void | setDxyRecorders (age_t AGE, bool patchwise) |
| |
|
| void | setAdultAlleleFreq () |
| |
| void | setOffspringAlleleFreq () |
| |
| void | setHeterozygosity (age_t AGE) |
| |
| void | setAdultHeterozygosity () |
| |
| void | setOffspringHeterozygosity () |
| |
| double | getGlobalAlleleFreq (unsigned int loc, unsigned int all) |
| |
| double | getHeterozygosity (unsigned int loc) |
| |
|
| void | setAlleleTables (age_t AGE) |
| |
| void | setHeteroTable (age_t AGE) |
| |
| void | allocateTables (unsigned int loci, unsigned int all) |
| |
| DataTable< double > * | getAlleleFreqTable () |
| | Accessor to the table of allele frequencies, per patch. More...
|
| |
| DataTable< unsigned int > * | getAlleleCountTable () |
| |
| DataTable< double > * | getHeteroTable () |
| |
| TMatrix * | getGlobalFreqs () |
| | Accessor to the table of allele frequencies in the whole population. More...
|
| |
| void | setFstMatrix (age_t AGE, unsigned char dim) |
| | Computes the weighted within and between patch Fst's as well as the overall Fst (Theta). More...
|
| |
| void | setAdultsFstMatrix () |
| |
| void | setAdultsFstWithin () |
| |
| void | setAdultsFstBetween () |
| |
| void | setOffsprgFstMatrix () |
| |
| void | setOffsprgFstWithin () |
| |
| void | setOffsprgFstBetween () |
| |
| double | getWeightedFst () |
| | Returns the weighted Fst using Weir & Hill (2002) method. More...
|
| |
| double | getFst_ij (unsigned int i) |
| | Accessor to the Fst matrix as set by setFstMatrix(). More...
|
| |
| void | setFst_li (unsigned int N, unsigned int L, double **array) |
| | Computes the per-locus per-patch Fst values using Weir&Hill 2002 approach. More...
|
| |
| void | setFstat (age_t AGE) |
| | Computes the F-statistics following Nei & Chesser (1983). More...
|
| |
| void | setFstat_bitstring (age_t AGE) |
| | Streaming F-stat computation for diallelic bitstring traits. More...
|
| |
| void | setOffsprgFstat () |
| |
| void | setAdultsFstat () |
| |
| double | setHo (age_idx age_pos) |
| |
| double | setHs (age_idx age_pos) |
| |
| double | setHt (age_idx age_pos) |
| |
| double | getHsnei () |
| |
| double | getHtnei () |
| |
| double | getHo () |
| |
| double | getHs () |
| |
| double | getHt () |
| |
| double | getFst () |
| |
| double | getFis () |
| |
| double | getFit () |
| |
| deque< double > | setHo2 (age_idx age_pos) |
| | New version of Nei & Chesser. More...
|
| |
| deque< double > | setHs2 (age_idx age_pos) |
| |
| deque< double > | setHt2 (age_idx age_pos) |
| |
| void | setFstatWeirCockerham (age_t AGE) |
| | Computes the Weir & Cockerham (1984) Fstat values (Theta, F, and f). More...
|
| |
| void | setFstatWeirCockerham_bitstring (age_t AGE) |
| | Streaming W&C Fstat for diallelic bitstring traits. More...
|
| |
| void | setFstatWeirCockerham_MS (age_t AGE) |
| |
| void | setOffspringFstatWeirCockerham () |
| |
| void | setAdultsFstatWeirCockerham () |
| |
| double | getFstWC () |
| |
| double | getFisWC () |
| |
| double | getFitWC () |
| |
| void | setLociDivCounter (age_t AGE) |
| | Sets the allelic diversity counters. More...
|
| |
| double | getNbAllLocal () |
| |
| double | getNbAllGlobal () |
| |
| double | getFixLocLocal () |
| |
| double | getFixLocGlobal () |
| |
|
| double | Coancestry (const TTrait *ind1, const TTrait *ind2, unsigned int nb_locus) |
| | Gives the coancestry (probability of identity by state) of two gene sequences. More...
|
| |
| void | setCoaMatrix (age_idx age_pos, unsigned char dim) |
| | Computes the within and between patches coancestry coefficients. More...
|
| |
| void | setAdultsCoaMatrix () |
| |
| void | setOffsprgCoaMatrix () |
| |
| void | setAdultsCoaWithin () |
| |
| void | setOffsprgCoaWithin () |
| |
| void | setAdultsCoaBetween () |
| |
| void | setOffsprgCoaBetween () |
| |
| void | setAdults_Theta () |
| |
| double | getCoa (unsigned int i) |
| | Gets the given coancestry coefficient from the coancestry matrix. More...
|
| |
| double | getMeanTheta () |
| |
| double | getMeanAlpha () |
| |
| double | getTheta_FF () |
| | Gives the mean within females coancestry coefficient. More...
|
| |
| double | getTheta_MM () |
| | Gives the mean within males coancestry coefficient. More...
|
| |
| double | getTheta_FM () |
| | Gives the mean between males and females coancestry coefficient. More...
|
| |
| void | setSibStats () |
| |
| void | setSibCoa (Individual *I1, Individual *I2) |
| |
| double | getSibProportions (unsigned int i) |
| |
| double | getSibCoaMeans (unsigned int i) |
| |
|
| void | setAdltNeiGeneticDistance () |
| |
| void | setOffsprgNeiGeneticDistance () |
| |
| void | setNeiGeneticDistance (age_t AGE) |
| |
| double | getNeiGeneticDistance (unsigned int i) |
| |
| double | getMeanNeiGeneticDistance () |
| |
|
| double | getDxyOffspringPerPatch (unsigned int patch1, unsigned patch2) |
| |
| double | getDxyAdultPerPatch (unsigned int patch1, unsigned patch2) |
| |
| double | getDxyPerPatch (age_idx age, unsigned int patch1, unsigned patch2) |
| |
| double | getDxy (unsigned int age_class) |
| |
 Public Member Functions inherited from TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH > Public Member Functions inherited from TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH > |
| | TraitStatHandler (TProtoNeutralGenes *trait_proto) |
| |
| virtual | ~TraitStatHandler () |
| |
 Public Member Functions inherited from StatHandler< SH > Public Member Functions inherited from StatHandler< SH > |
| | StatHandler () |
| |
| virtual | ~StatHandler () |
| |
| virtual void | clear () |
| | Empties the _recorders list, they are destroyed in StatHandlerBase::reset(). More...
|
| |
| virtual StatRecorder< SH > * | add (std::string Title, std::string Name, age_t AGE, unsigned int ARG1, unsigned int ARG2, double(SH::*getStatNoArg)(void), double(SH::*getStatOneArg)(unsigned int), double(SH::*getStatTwoArg)(unsigned int, unsigned int), void(SH::*setStat)(void)) |
| | Adds a StatRecorder to the list, it is also added to the StatHandlerBase::_stats list. More...
|
| |
 Public Member Functions inherited from StatHandlerBase Public Member Functions inherited from StatHandlerBase |
| | StatHandlerBase () |
| |
| virtual | ~StatHandlerBase () |
| |
| virtual void | reset () |
| | Empties the _stats list and calls clear() (defined in the derived class). More...
|
| |
| Metapop * | get_pop_ptr () |
| |
| void | set_service (StatServices *srv) |
| |
| StatServices * | get_service () |
| |
| unsigned int | getOccurrence () |
| |
| unsigned int | getNumOccurrences () |
| |
| unsigned int | getCurrentOccurrence () |
| |
| unsigned int | getNbRecorders () |
| |
| std::list< StatRecBase * > & | getStats () |
| |
| virtual void | add (StatRecBase *rec) |
| |
| virtual void | update () |
| | This function is left empty as the StatServices calls StatRecorder::setVal directly. More...
|
| |
 Public Member Functions inherited from Handler Public Member Functions inherited from Handler |
| virtual | ~Handler () |
| |
The stat handler for neutral markers.
| void TTNeutralGenesSH::setAlleleTables |
( |
age_t |
AGE | ) |
|
97 unsigned int* count1 =
new unsigned int[nb_locus]();
99 for(
unsigned int k = 0; k < patchNbr; ++k) {
102 memset(count1, 0, nb_locus *
sizeof(
unsigned int));
104 for (
int sx = 0; sx < 2; ++sx) {
106 for(
unsigned int i = 0, size = crnt_patch->
size(
sex_t(sx), age_pos); i < size; ++i) {
111 for (
int chr = 0; chr < 2; ++chr) {
114 for (
size_t w = 0; w < nwords; ++w) {
125 for (
unsigned int j = 0; j < nb_locus; ++j) {
128 crnt_patch->
size(age_pos) * 2 - count1[j]);
137 for(
unsigned int k = 0; k < patchNbr; ++k) {
141 for (
int sx = 0; sx < 2; ++sx) {
143 for(
unsigned int i = 0, size = crnt_patch->
size(
sex_t(sx), age_pos); i < size; ++i) {
147 for (
unsigned int j = 0; j < nb_locus; ++j)
150 for (
unsigned int j = 0; j < nb_locus; ++j)
159 for (
unsigned int i = 0; i < patchNbr; ++i) {
165 for (
unsigned int l = 0; l < nb_locus; ++l)
166 for (
unsigned int u = 0; u < nb_allele; ++u)
170 for (
unsigned int l = 0; l < nb_locus; ++l)
171 for (
unsigned int u = 0; u < nb_allele; ++u)
177 unsigned int tot_size =
_pop->
size(AGE) * 2;
181 for(
unsigned int i = 0; i < patchNbr; i++)
182 for (
unsigned int l = 0; l < nb_locus; ++l)
183 for (
unsigned int u = 0; u < nb_allele; ++u)
186 for (
unsigned int l = 0; l < nb_locus; ++l)
187 for (
unsigned int u = 0; u < nb_allele; ++u)
#define BITS_PER_WORD
Definition: bitstring.h:40
void set(unsigned int group, unsigned int Class, unsigned int elmnt, T val)
Sets the element 'elmnt' of the class 'Class' in the group 'group' to the value 'val'.
Definition: datatable.h:231
void init(T val)
Sets all elements of the table to value 'val'.
Definition: datatable.h:255
void increment(unsigned int group, unsigned int Class, unsigned int elmnt)
Increments 'elmnt' of the class 'Class' in the group 'group' by one.
Definition: datatable.h:235
void assign(double val)
Assigns a value to all element of the matrix.
Definition: tmatrix.h:154
void divide(unsigned int i, unsigned int j, double value)
Divide an element of the matrix by a value.
Definition: tmatrix.h:315
void plus(unsigned int i, unsigned int j, double value)
Adds a value to an element of the matrix.
Definition: tmatrix.h:255
TTNeutralGenes_bitstring : diallelic neutral loci encoded as bitstrings.
Definition: ttneutralgenes_bitstring.h:39
const bitstring & get_bit_sequence(bool chromosome) const
Definition: ttneutralgenes_bitstring.h:51
Non-template and faster implementation of std::bitset.
Definition: bitstring.h:56
unsigned long _ul
Definition: bitstring.h:60
_ul * getword_atIdx(size_t index) const
Definition: bitstring.h:158
size_t nb_words() const
Definition: bitstring.h:163
sex_t
Sex types, males are always 0 and females 1!!
Definition: types.h:35
References _alleleCountTable, _alleleFreqTable, _globalAlleleFreq, _is_diallelic_bitstring, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTraitIndex, _table_set_age, _table_set_gen, _table_set_repl, ADLTx, ADULTS, allocateTables(), TMatrix::assign(), BITS_PER_WORD, TMatrix::divide(), Patch::get(), DataTable< T >::get(), TTrait::get_allele(), TProtoNeutralGenes::get_allele_num(), TTNeutralGenes_bitstring::get_bit_sequence(), TProtoNeutralGenes::get_locus_num(), Metapop::getCurrentGeneration(), Metapop::getCurrentReplicate(), DataTable< T >::getNumGroups(), Metapop::getPatch(), Metapop::getPatchNbr(), Individual::getTrait(), bitstring::getword_atIdx(), DataTable< T >::increment(), DataTable< T >::init(), bitstring::nb_words(), OFFSx, TMatrix::plus(), DataTable< T >::set(), Metapop::size(), and Patch::size().
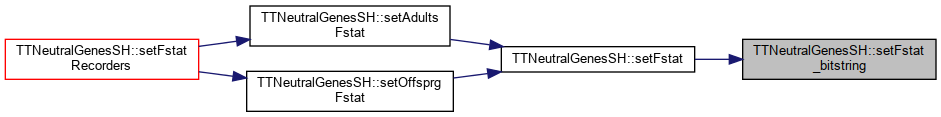
Referenced by setAdultAlleleFreq(), setFst_li(), setFstat(), setFstatWeirCockerham(), setFstatWeirCockerham_MS(), setFstMatrix(), setNeiGeneticDistance(), setOffspringAlleleFreq(), and TTNeutralGenesFH::write_varcompWC().
| void TTNeutralGenesSH::setFstat |
( |
age_t |
AGE | ) |
|
Computes the F-statistics following Nei & Chesser (1983).
288 double harmonic = 0, nbpatch = 0, nbind;
301 for (
unsigned int i = 0; i < patchNbr; ++i){
306 harmonic += 1.0/nbind;
310 harmonic = nbpatch / harmonic;
316 _hsnei = (nbpatch != 0 ? harmonic/(harmonic-1.0)*(
_hs-(
_ho/(2.0*harmonic))) : nanf(
"NULL") );
318 -(
_ho/(2.0*harmonic*nbpatch)) : nanf(
"NULL") );
double setHo(age_idx age_pos)
Definition: stats_fstat.cc:534
double setHt(age_idx age_pos)
Definition: stats_fstat.cc:639
void setLociDivCounter(age_t AGE)
Sets the allelic diversity counters.
Definition: stats_fstat.cc:466
void setFstat_bitstring(age_t AGE)
Streaming F-stat computation for diallelic bitstring traits.
Definition: stats_fstat.cc:327
double setHs(age_idx age_pos)
Definition: stats_fstat.cc:594
References _alleleCountTable, _fis, _fit, _fst, _ho, _hs, _hsnei, _ht, _htnei, _is_diallelic_bitstring, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, ADLTx, ADULTS, allocateTables(), TProtoNeutralGenes::get_allele_num(), TProtoNeutralGenes::get_locus_num(), DataTable< T >::getNumGroups(), Metapop::getPatchNbr(), OFFSx, setAlleleTables(), setFstat_bitstring(), setHo(), setHs(), setHt(), setLociDivCounter(), and Metapop::size().
Referenced by setAdultsFstat(), and setOffsprgFstat().
| void TTNeutralGenesSH::setFstat_bitstring |
( |
age_t |
AGE | ) |
|
Streaming F-stat computation for diallelic bitstring traits.
Processes loci in word-sized chunks (64 loci) to avoid allocating per-patch x per-locus DataTables.
334 double harmonic = 0, nbpatch = 0;
335 unsigned int* patch_2N =
new unsigned int[patchNbr];
337 for (
unsigned int i = 0; i < patchNbr; ++i) {
338 unsigned int psize =
_pop->
size(AGE, i);
339 patch_2N[i] = psize * 2;
342 harmonic += 1.0 / psize;
345 harmonic = (nbpatch > 0) ? nbpatch / harmonic : 0;
347 unsigned int total_2N =
_pop->
size(AGE) * 2;
356 double hs_sum = 0, ht_sum = 0;
357 unsigned int nb_all_local_sum = 0;
358 unsigned int fix_loc_local_count = 0, fix_loc_global_count = 0;
359 bool pop_has_allele1, pop_has_allele0;
360 double nb_all_global_sum = 0;
363 unsigned int* count1 =
new unsigned int[patchNbr *
BITS_PER_WORD];
365 for (
size_t w = 0; w < nwords; ++w) {
371 memset(count1, 0, patchNbr *
BITS_PER_WORD *
sizeof(
unsigned int));
374 for (
unsigned int k = 0; k < patchNbr; ++k) {
378 for (
int sx = 0; sx < 2; ++sx) {
379 for (
unsigned int ind = 0, sz = patch->
size(
sex_t(sx), age_pos);
386 for (
int chr = 0; chr < 2; ++chr) {
389 count1[base + __builtin_ctzl(word)]++;
398 for (
unsigned int bit = 0; bit < loci_in_word; ++bit) {
400 unsigned int global_count1 = 0;
401 pop_has_allele1 =
false;
402 pop_has_allele0 =
false;
404 for (
unsigned int k = 0; k < patchNbr; ++k) {
405 if (patch_2N[k] == 0)
continue;
408 unsigned int c0 = patch_2N[k] - c1;
412 double p = (double)c1 / patch_2N[k];
413 hs_sum += 2.0 * p * (1.0 - p);
416 nb_all_local_sum += (c0 > 0) + (c1 > 0);
419 fix_loc_local_count += (c1 == 0 || c1 == patch_2N[k]);
421 pop_has_allele1 |= (c1 > 0);
422 pop_has_allele0 |= (c0 > 0);
427 double p_global = (double)global_count1 / total_2N;
428 ht_sum += 2.0 * p_global * (1.0 - p_global);
432 nb_all_global_sum += (pop_has_allele0 ? 1.0 : 0.0)
433 + (pop_has_allele1 ? 1.0 : 0.0);
436 fix_loc_global_count += (global_count1 == 0 || global_count1 == total_2N);
443 _hs = (nbpatch > 0) ? hs_sum / (nloc * nbpatch) : 0;
444 _ht = (nloc > 0) ? ht_sum / nloc : 0;
446 _nb_all_local = (nbpatch > 0) ? nb_all_local_sum / (
double)(nloc * nbpatch)
449 _fix_loc_local = (nbpatch > 0) ? (
double)fix_loc_local_count / nbpatch : 0;
453 _hsnei = (nbpatch > 0 ? harmonic/(harmonic-1.0)*(
_hs-(
_ho/(2.0*harmonic)))
456 -(
_ho/(2.0*harmonic*nbpatch))
#define BITSET_WORDS(__n)
Definition: bitstring.h:44
References _fis, _fit, _fix_loc_global, _fix_loc_local, _fst, _ho, _hs, _hsnei, _ht, _htnei, _nb_all_global, _nb_all_local, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTraitIndex, ADLTx, ADULTS, BITS_PER_WORD, BITSET_WORDS, Patch::get(), TTNeutralGenes_bitstring::get_bit_sequence(), TProtoNeutralGenes::get_locus_num(), Metapop::getPatch(), Metapop::getPatchNbr(), Individual::getTrait(), bitstring::getword_atIdx(), OFFSx, setHo(), Metapop::size(), and Patch::size().
Referenced by setFstat().
| void TTNeutralGenesSH::setFstatWeirCockerham |
( |
age_t |
AGE | ) |
|
Computes the Weir & Cockerham (1984) Fstat values (Theta, F, and f).
990 double *pop_sizes =
new double [patchNbr];
991 double tot_size, inv_ntot;
992 double sum_weights = 0;
993 double nbar, nc, inv_nbar;
994 unsigned int extantPs = 0;
998 for(
unsigned int i = 0; i < patchNbr; i++) {
1002 sum_weights += (pop_sizes[i] * pop_sizes[i] / tot_size);
1005 nbar = tot_size/extantPs;
1006 nc = (tot_size - sum_weights)/(extantPs-1);
1007 inv_nbar = 1/(nbar - 1);
1008 inv_ntot = 1/tot_size;
1011 double s2, pbar, hbar;
1012 double s2_denom = 1.0/((extantPs-1)*nbar),
1013 r = (double)(extantPs-1)/extantPs,
1014 hbar_factor=(2*nbar-1)/(4*nbar);
1015 double a = 0, b = 0, c = 0, x;
1017 for (
unsigned int l = 0; l < nb_locus; ++l) {
1019 for (
unsigned int u = 0; u < nb_allele; ++u) {
1021 s2 = pbar = hbar = 0;
1023 for (
unsigned int i = 0; i < patchNbr; ++i) {
1029 s2 += var * pop_sizes[i];
1039 x = pbar * (1 - pbar) - r * s2;
1040 a += s2 - inv_nbar*( x - 0.25 * hbar);
1041 b += x - hbar_factor * hbar;
1050 b *= nbar/(nbar - 1);
1054 _fit_WC = (a + b) / (a + b + c);
1057 delete [] pop_sizes;
void setFstatWeirCockerham_bitstring(age_t AGE)
Streaming W&C Fstat for diallelic bitstring traits.
Definition: stats_fstat.cc:1062
void setHeteroTable(age_t AGE)
Definition: stats_fstat.cc:199
References _alleleFreqTable, _fis_WC, _fit_WC, _fst_WC, _globalAlleleFreq, _heteroTable, _is_diallelic_bitstring, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, DataTable< T >::get(), TMatrix::get(), TProtoNeutralGenes::get_allele_num(), TProtoNeutralGenes::get_locus_num(), Metapop::getPatchNbr(), setAlleleTables(), setFstatWeirCockerham_bitstring(), setHeteroTable(), and Metapop::size().
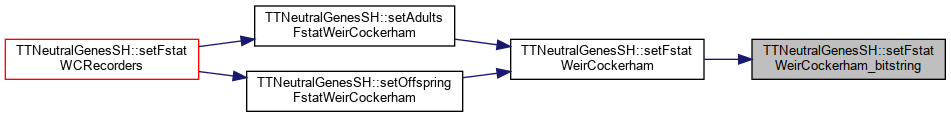
Referenced by setAdultsFstatWeirCockerham(), and setOffspringFstatWeirCockerham().
| void TTNeutralGenesSH::setFstatWeirCockerham_bitstring |
( |
age_t |
AGE | ) |
|
Streaming W&C Fstat for diallelic bitstring traits.
1069 double *pop_sizes =
new double[patchNbr];
1071 double inv_ntot = 1.0 / tot_size;
1072 double sum_weights = 0;
1073 unsigned int extantPs = 0;
1075 for (
unsigned int i = 0; i < patchNbr; i++) {
1079 sum_weights += (pop_sizes[i] * pop_sizes[i] / tot_size);
1083 double nbar = tot_size / extantPs;
1084 double nc = (tot_size - sum_weights) / (extantPs - 1);
1085 double inv_nbar = 1.0 / (nbar - 1);
1086 double s2_denom = 1.0 / ((extantPs - 1) * nbar);
1087 double r = (double)(extantPs - 1) / extantPs;
1088 double hbar_factor = (2 * nbar - 1) / (4 * nbar);
1090 double a = 0, b = 0, c = 0;
1095 unsigned int* count1 =
new unsigned int[patchNbr *
BITS_PER_WORD];
1096 unsigned int* het =
new unsigned int[patchNbr *
BITS_PER_WORD];
1098 for (
size_t w = 0; w < nwords; ++w) {
1104 memset(count1, 0, patchNbr *
BITS_PER_WORD *
sizeof(
unsigned int));
1105 memset(het, 0, patchNbr *
BITS_PER_WORD *
sizeof(
unsigned int));
1107 for (
unsigned int k = 0; k < patchNbr; ++k) {
1111 for (
int sx = 0; sx < 2; ++sx) {
1112 for (
unsigned int ind = 0, sz = patch->
size(
sex_t(sx), age_pos);
1124 while (word) { count1[base + __builtin_ctzl(word)]++; word &= word - 1; }
1126 while (word) { count1[base + __builtin_ctzl(word)]++; word &= word - 1; }
1130 while (word) { het[base + __builtin_ctzl(word)]++; word &= word - 1; }
1138 for (
unsigned int bit = 0; bit < loci_in_word; ++bit) {
1140 unsigned int global_count1 = 0;
1143 for (
unsigned int k = 0; k < patchNbr; ++k) {
1144 if (pop_sizes[k] == 0)
continue;
1149 double pbar = (double)global_count1 / (2.0 * tot_size);
1153 for (
unsigned int k = 0; k < patchNbr; ++k) {
1154 if (pop_sizes[k] == 0)
continue;
1155 double p_i = (double)count1[k *
BITS_PER_WORD + bit] / (2.0 * pop_sizes[k]);
1156 double var = p_i - pbar;
1157 s2 += var * var * pop_sizes[k];
1161 double x = pbar * (1 - pbar) - r * s2;
1162 a += s2 - inv_nbar * (x - 0.25 * hbar);
1163 b += x - hbar_factor * hbar;
1169 b *= nbar / (nbar - 1);
1173 _fit_WC = (a + b) / (a + b + c);
References _fis_WC, _fit_WC, _fst_WC, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTraitIndex, ADLTx, ADULTS, BITS_PER_WORD, BITSET_WORDS, Patch::get(), TTNeutralGenes_bitstring::get_bit_sequence(), TProtoNeutralGenes::get_locus_num(), Metapop::getPatch(), Metapop::getPatchNbr(), Individual::getTrait(), bitstring::getword_atIdx(), OFFSx, Metapop::size(), and Patch::size().
Referenced by setFstatWeirCockerham().
| void TTNeutralGenesSH::setFstatWeirCockerham_MS |
( |
age_t |
AGE | ) |
|
Computes W&C F-stats using the Mean Squares approach, similar to the implementation in Hierfstat. This code gives the exact same results as the other version.
1196 double *pop_sizes =
new double [patchNbr];
1198 double sum_weights = 0;
1200 unsigned int extantPs = 0;
1204 for(
unsigned int i = 0; i < patchNbr; i++) {
1208 sum_weights += (pop_sizes[i] * pop_sizes[i] / tot_size);
1212 nc = (tot_size - sum_weights)/(extantPs-1);
1215 unsigned int npl = extantPs;
1220 unsigned int *alploc =
new unsigned int [nb_locus];
1222 unsigned int **alploc_table =
new unsigned int* [nb_locus];
1224 for(
unsigned int i = 0; i < nb_locus; ++i)
1225 alploc_table[i] =
new unsigned int[nb_allele];
1227 unsigned int tot_num_allele = 0;
1229 for(
unsigned int l = 0; l < nb_locus; ++l){
1233 for(
unsigned int cnt, a = 0; a < nb_allele; ++a) {
1237 for(
unsigned int i = 0; i < patchNbr; i++) {
1242 alploc_table[l][a] = (cnt != 0);
1243 alploc[l] += (cnt != 0);
1246 tot_num_allele += alploc[l];
1254 double *SSG =
new double[tot_num_allele];
1255 double *SSP =
new double[tot_num_allele];
1256 double *SSi =
new double[tot_num_allele];
1258 unsigned int all_cntr = 0;
1260 double het, freq, var;
1262 for(
unsigned int l = 0; l < nb_locus; ++l) {
1264 for(
unsigned int a = 0; a < nb_allele & all_cntr < tot_num_allele; ++a) {
1266 if(alploc_table[l][a] == 0)
continue;
1272 for(
unsigned int p = 0; p < patchNbr; ++p){
1284 SSG[all_cntr] += het;
1286 SSi[all_cntr] += 2*pop_sizes[p]*freq*(1-freq) - het/2;
1288 SSP[all_cntr] += 2*pop_sizes[p]*var;
1297 assert(all_cntr == tot_num_allele);
1299 double *MSG =
new double[tot_num_allele];
1300 double *MSP =
new double[tot_num_allele];
1301 double *MSI =
new double[tot_num_allele];
1302 double *sigw =
new double[tot_num_allele];
1303 double *siga =
new double[tot_num_allele];
1304 double *sigb =
new double[tot_num_allele];
1309 double SIGA = 0, SIGB = 0, SIGW = 0;
1311 for(
unsigned int i = 0; i < tot_num_allele; ++i){
1313 MSG[i] = SSG[i] / (2 * tot_size);
1316 MSP[i] = SSP[i] / (npl-1);
1318 MSI[i] = SSi[i]/ (tot_size - npl);
1320 sigb[i] = 0.5*(MSI[i] - MSG[i]);
1322 siga[i] = (MSP[i] - MSI[i])/(2*nc);
1337 double lsiga, lsigb, lsigw;
1341 for(
unsigned int allcntr = 0, i = 0; i < nb_locus; ++i) {
1343 lsiga = lsigb = lsigw = 0;
1345 for(
unsigned int l = 0; l < alploc[i]; ++l) {
1347 lsiga += siga[allcntr];
1348 lsigb += sigb[allcntr];
1349 lsigw += sigw[allcntr];
1357 _fit_WC_loc[i] = (lsiga +lsigb) /(lsiga + lsigb + lsigw);
1363 _fst_WC = SIGA / (SIGA + SIGB + SIGW);
1364 _fit_WC = (SIGA + SIGB) / (SIGA + SIGB + SIGW);
1365 _fis_WC = SIGB / (SIGB + SIGW);
1369 for(
unsigned int i = 0; i < nb_locus; ++i)
1370 delete[]alploc_table[i];
1371 delete[]alploc_table;
References _alleleCountTable, _alleleFreqTable, _fis_WC, _fis_WC_loc, _fit_WC, _fit_WC_loc, _fst_WC, _fst_WC_loc, _globalAlleleFreq, _heteroTable, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, DataTable< T >::get(), TMatrix::get(), TProtoNeutralGenes::get_allele_num(), TProtoNeutralGenes::get_locus_num(), Metapop::getPatchNbr(), setAlleleTables(), setHeteroTable(), and Metapop::size().
| void TTNeutralGenesSH::setFstMatrix |
( |
age_t |
AGE, |
|
|
unsigned char |
dim |
|
) |
| |
Computes the weighted within and between patch Fst's as well as the overall Fst (Theta).
The method used here is that of Weir & Hill 2002, Ann. Rev. Genet. 36:721-750. The weighting is done for samples (patches) of unequal sizes.
- Parameters
-
| AGE | the age class |
| dim | the dimension of the matrix to fill:
- 1 = the diagonal (i.e. the wihtin patch Fst or theta_ii)
- 2 = the upper half (i.e. the between patch Fst or theta_ii')
- 3 = both
|
888 double *pop_weights =
new double[patchNbr];
889 double *pop_sizes =
new double[patchNbr];
890 double **numerator =
new double*[patchNbr];
891 for(
unsigned int i = 0; i < patchNbr; i++) numerator[i] =
new double [patchNbr];
893 double numerator_W = 0;
894 double denominator = 0;
895 double sum_weights = 0;
899 for(
unsigned int i = 0; i < patchNbr; ++i) {
900 pop_sizes[i] =
_pop->
size(AGE, i) * 2;
901 pop_weights[i] = pop_sizes[i] - (pop_sizes[i] * pop_sizes[i] / tot_size);
902 sum_weights += pop_weights[i];
903 for(
unsigned int j = 0; j < patchNbr; j++)
907 double p, pq, var, num;
909 for (
unsigned int i = 0; i < patchNbr; ++i) {
911 if( !pop_sizes[i] )
continue;
913 for (
unsigned int l = 0; l < nb_locus; ++l) {
915 for (
unsigned int u = 0; u < nb_allele; ++u) {
925 num = pq * pop_sizes[i] / (pop_sizes[i] -1);
927 numerator[i][i] += num;
929 numerator_W += num * pop_sizes[i];
931 denominator += pop_sizes[i] * var + pop_weights[i] * pq;
937 for (
unsigned int i = 0; i < patchNbr; ++i) {
938 if( !pop_sizes[i] )
continue;
939 _fst_matrix->
set(i, i, 1 - (numerator[i][i] * sum_weights / denominator) );
941 _fst_WH = 1 - ((numerator_W * sum_weights) / (denominator * tot_size));
946 for (
unsigned int l = 0; l < nb_locus; ++l)
947 for (
unsigned int u = 0; u < nb_allele; ++u)
948 for (
unsigned int i = 0; i < patchNbr - 1; ++i) {
949 if( !pop_sizes[i] )
continue;
950 for (
unsigned int j = i + 1; j < patchNbr; ++j) {
951 if( !pop_sizes[j] )
continue;
954 numerator[i][j] += pi * (1 - pj) + pj * (1 - pi);
957 for (
unsigned int i = 0; i < patchNbr - 1; ++i){
958 if( !pop_sizes[i] )
continue;
959 for (
unsigned int j = i + 1; j < patchNbr; ++j){
960 if( !pop_sizes[j] )
continue;
961 _fst_matrix->
set(i, j, 1 - ( (numerator[i][j] * sum_weights) / (2 * denominator)) );
967 delete [] pop_weights;
969 for(
unsigned int i = 0; i < patchNbr; i++)
delete [] numerator[i];
void set(unsigned int i, unsigned int j, double val)
Sets element at row i and column j to value val.
Definition: tmatrix.h:102
References _alleleFreqTable, _fst_matrix, _fst_WH, _globalAlleleFreq, StatHandlerBase::_pop, TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >::_SHLinkedTrait, TMatrix::assign(), DataTable< T >::get(), TMatrix::get(), TProtoNeutralGenes::get_allele_num(), TProtoNeutralGenes::get_locus_num(), Metapop::getPatchNbr(), TMatrix::length(), TMatrix::reset(), TMatrix::set(), setAlleleTables(), and Metapop::size().
Referenced by setAdultsFstBetween(), setAdultsFstMatrix(), setAdultsFstWithin(), setOffsprgFstBetween(), setOffsprgFstMatrix(), and setOffsprgFstWithin().
 Inheritance diagram for TTNeutralGenesSH:
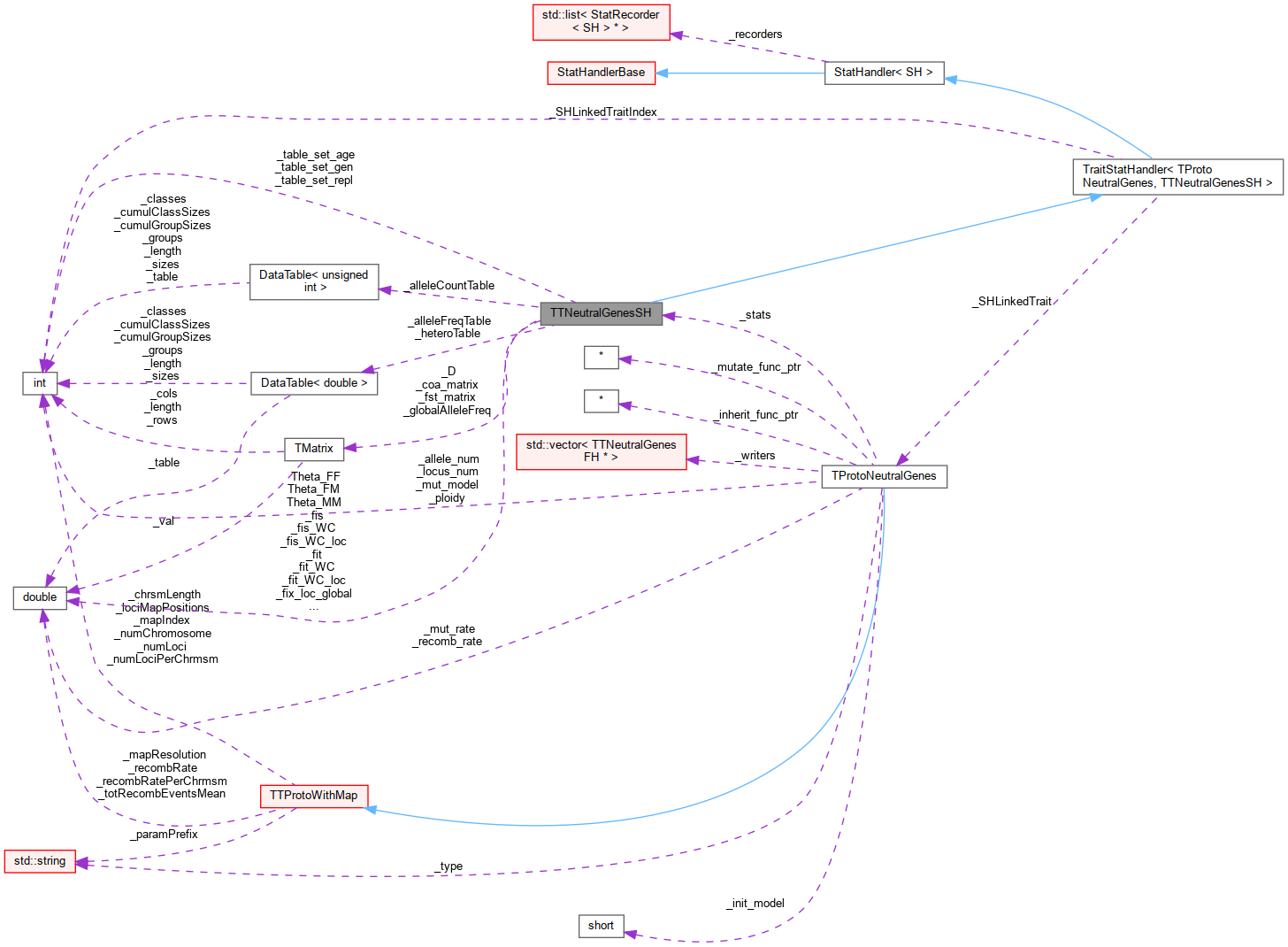
Inheritance diagram for TTNeutralGenesSH: Collaboration diagram for TTNeutralGenesSH:
Collaboration diagram for TTNeutralGenesSH: Public Member Functions inherited from TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >
Public Member Functions inherited from TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH > Public Member Functions inherited from StatHandler< SH >
Public Member Functions inherited from StatHandler< SH > Public Member Functions inherited from StatHandlerBase
Public Member Functions inherited from StatHandlerBase Public Member Functions inherited from Handler
Public Member Functions inherited from Handler Protected Types inherited from StatHandler< SH >
Protected Types inherited from StatHandler< SH > Protected Attributes inherited from TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH >
Protected Attributes inherited from TraitStatHandler< TProtoNeutralGenes, TTNeutralGenesSH > Protected Attributes inherited from StatHandler< SH >
Protected Attributes inherited from StatHandler< SH > Protected Attributes inherited from StatHandlerBase
Protected Attributes inherited from StatHandlerBase Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: Here is the caller graph for this function:
Here is the caller graph for this function: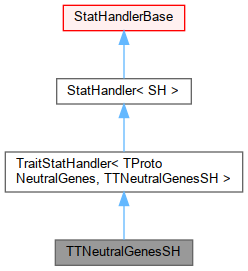


































































 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
