#include <ttquanti.h>
 Inheritance diagram for TTQuantiFH:
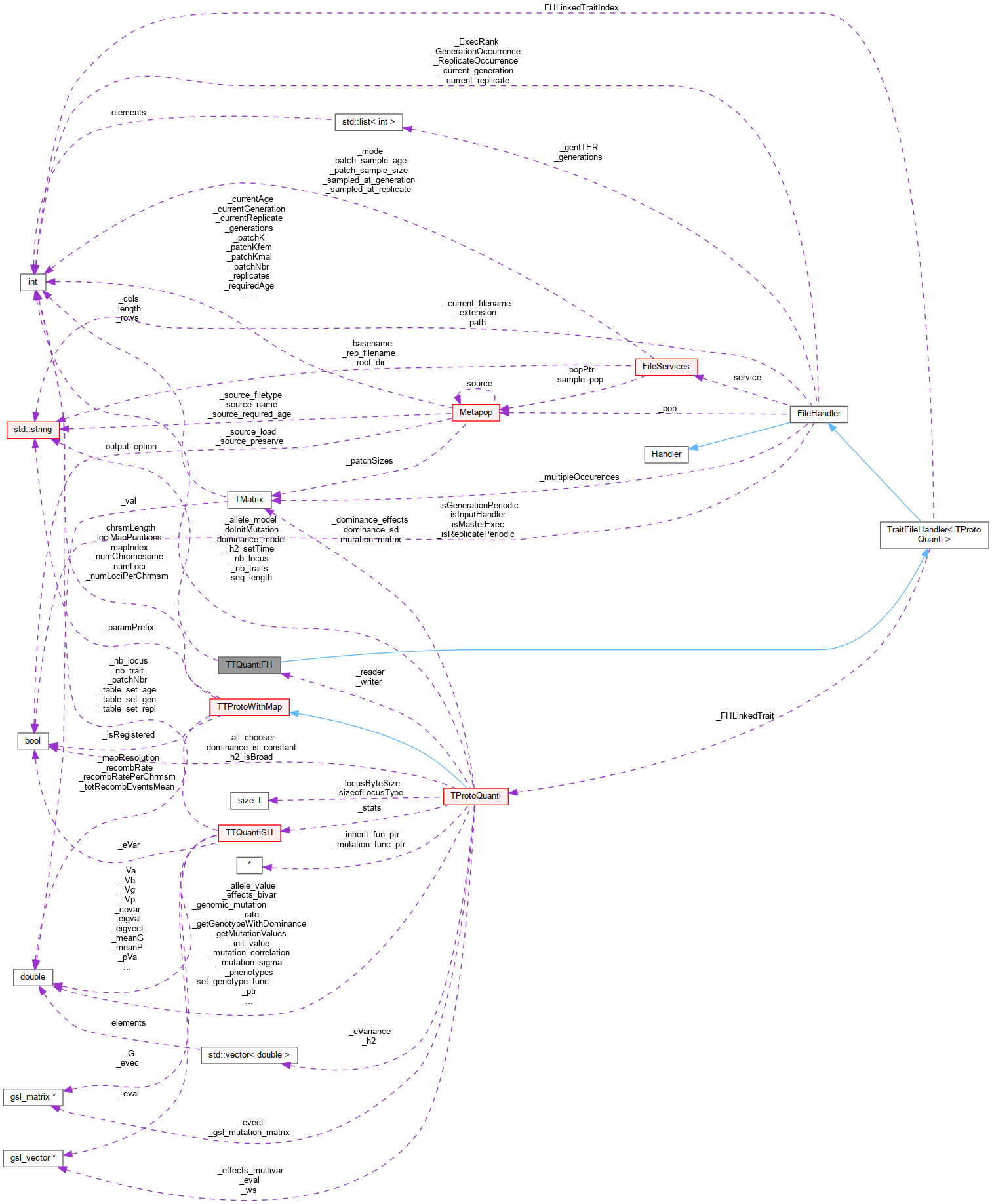
Inheritance diagram for TTQuantiFH: Collaboration diagram for TTQuantiFH:
Collaboration diagram for TTQuantiFH:Public Member Functions | |
| TTQuantiFH (TProtoQuanti *T) | |
| virtual | ~TTQuantiFH () |
| void | setOutputOption (string opt) |
| virtual void | FHwrite () |
| void | write_TABLE () |
| void | print (ofstream &FH, Patch *patch, sex_t SEX, age_idx Ax, unsigned int print_gene, bool print_genotype, bool print_additive_genotype) |
| void | write_PLINK () |
| void | print_PLINK_PED (ofstream &FH, age_idx Ax, Patch *patch) |
| void | print_PLINK_FAM (ofstream &FH, age_idx Ax, Patch *patch) |
| virtual void | FHread (string &filename) |
 Public Member Functions inherited from TraitFileHandler< TProtoQuanti > Public Member Functions inherited from TraitFileHandler< TProtoQuanti > | |
| TraitFileHandler (TProtoQuanti *trait_proto, const char *ext) | |
| virtual | ~TraitFileHandler () |
| virtual void | FHread (string &filename)=0 |
| virtual void | set (bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path, TProtoQuanti *trait_proto) |
| virtual void | set_multi (bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path, TProtoQuanti *trait_proto) |
 Public Member Functions inherited from FileHandler Public Member Functions inherited from FileHandler | |
| FileHandler (const char *ext) | |
| virtual | ~FileHandler () |
| virtual void | init () |
| Called by notifier during simulation setup, performs file checking. More... | |
| virtual vector< string > | ifExist () |
| Checks if any file associated with the current file name already exists on disk. More... | |
| virtual void | set (bool rpl_per, bool gen_per, int rpl_occ, int gen_occ, int rank, string path) |
| Sets the hanlder parameters. More... | |
| virtual void | set_multi (bool rpl_per, bool gen_per, int rpl_occ, TMatrix *Occ, string path) |
| virtual void | update () |
| Updates the inner replicate and generation counters and calls FHwrite if needed by the the periodicity of the file. More... | |
| Metapop * | get_pop_ptr () |
| Returns the pointer to the current metapop through the FileServices interface. More... | |
| void | set_pop_ptr (Metapop *pop_ptr) |
| FileServices * | get_service () |
| Returns pointer to the FileServices. More... | |
| void | set_service (FileServices *srv) |
| std::string & | get_path () |
| void | set_path () |
| std::string & | get_extension () |
| void | set_extension (const char *ext) |
| std::string & | get_filename () |
| Builds and returns the current file name depending on the periodicity of the file. More... | |
| bool | get_isInputHandler () |
| void | set_isInputHandler (bool val) |
| bool | get_isReplicatePeriodic () |
| void | set_isReplicatePeriodic (bool val) |
| unsigned int | get_ReplicateOccurrence () |
| void | set_ReplicateOccurrence (unsigned int val) |
| bool | get_isGenerationPeriodic () |
| void | set_isGenerationPeriodic (bool val) |
| unsigned int | get_GenerationOccurrence () |
| void | set_GenerationOccurrence (unsigned int val) |
| unsigned int | get_ExecRank () |
| unused yet... More... | |
| void | set_ExecRank (int val) |
| TMatrix * | get_OccMatrix () |
| void | set_OccMatrix (TMatrix *occ) |
| bool | get_isMasterExec () |
| void | set_isMasterExec (bool is) |
 Public Member Functions inherited from Handler Public Member Functions inherited from Handler | |
| virtual | ~Handler () |
Private Attributes | |
| string | _output_option |
| bool | _has_genetic_map |
Additional Inherited Members | |
 Protected Attributes inherited from TraitFileHandler< TProtoQuanti > Protected Attributes inherited from TraitFileHandler< TProtoQuanti > | |
| TProtoQuanti * | _FHLinkedTrait |
| int | _FHLinkedTraitIndex |
 Protected Attributes inherited from FileHandler Protected Attributes inherited from FileHandler | |
| Metapop * | _pop |
| Pointer to the current metapop, set during initialization within the init function. More... | |
Detailed Description
Constructor & Destructor Documentation
◆ TTQuantiFH()
|
inline |
◆ ~TTQuantiFH()
Member Function Documentation
◆ FHread()
|
virtual |
Implements FileHandler.
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, TraitFileHandler< TProtoQuanti >::_FHLinkedTraitIndex, FileHandler::_pop, Patch::add(), error(), fatal(), TProtoQuanti::get_allele_model(), TProtoQuanti::get_dominance_model(), TProtoQuanti::get_env_var(), TProtoQuanti::get_locus_seq_pos(), TProtoQuanti::get_num_locus(), TProtoQuanti::get_num_traits(), TProtoQuanti::get_seq_diallele_value(), TProtoQuanti::get_seq_length(), Metapop::getPatch(), Metapop::getPatchNbr(), Individual::getTrait(), IndFactory::makeNewIndividual(), TTQuanti_continuous::set_sequence(), TTrait::set_sequence(), TTQuanti::set_value(), Individual::setAge(), and Individual::setPedigreeClass().
◆ FHwrite()
|
virtual |
Implements TraitFileHandler< TProtoQuanti >.
References _output_option, FileHandler::_pop, fatal(), FileServices::get_pop_ptr(), FileHandler::get_service(), FileServices::getSampledPop(), Metapop::isAlive(), write_PLINK(), and write_TABLE().
◆ print()
| void TTQuantiFH::print | ( | ofstream & | FH, |
| Patch * | patch, | ||
| sex_t | SEX, | ||
| age_idx | Ax, | ||
| unsigned int | print_gene, | ||
| bool | print_genotype, | ||
| bool | print_additive_genotype | ||
| ) |
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, TraitFileHandler< TProtoQuanti >::_FHLinkedTraitIndex, FEM, Patch::get(), TTQuanti::get_additive_genotype(), TTQuanti::get_allele_bit(), TTrait::get_allele_value(), TTQuanti::get_full_genotype(), TProtoQuanti::get_locus_seq_pos(), TProtoQuanti::get_num_locus(), TProtoQuanti::get_num_traits(), Individual::getFather(), Individual::getFatherID(), Individual::getHome(), Individual::getID(), Patch::getID(), Individual::getMother(), Individual::getMotherID(), Individual::getPedigreeClass(), Individual::getSex(), Individual::getTrait(), TTQuanti::getValue(), MAL, and Patch::size().
Referenced by write_TABLE().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ print_PLINK_FAM()
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, TraitFileHandler< TProtoQuanti >::_FHLinkedTraitIndex, FEM, Patch::get(), TProtoQuanti::get_num_traits(), Individual::getFatherID(), Individual::getHome(), Individual::getID(), Individual::getMotherID(), Individual::getTraitValue(), MAL, OFFSx, and Patch::size().
Referenced by write_PLINK().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ print_PLINK_PED()
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, TraitFileHandler< TProtoQuanti >::_FHLinkedTraitIndex, FEM, Patch::get(), TProtoQuanti::get_locus_seq_pos(), TProtoQuanti::get_num_locus(), TProtoQuanti::get_num_traits(), Individual::getFatherID(), Individual::getHome(), Individual::getID(), Individual::getMotherID(), Individual::getTrait(), Individual::getTraitValue(), MAL, OFFSx, and Patch::size().
Referenced by write_PLINK().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setOutputOption()
| void TTQuantiFH::setOutputOption | ( | string | opt | ) |
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, _has_genetic_map, TTProtoWithMap::_map, _output_option, GeneticMap::checkRegisteredTrait(), fatal(), TProtoQuanti::get_type(), and warning().
Referenced by TProtoQuanti::loadFileServices().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_PLINK()
| void TTQuantiFH::write_PLINK | ( | ) |
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, _has_genetic_map, TTProtoWithMap::_map, FileHandler::_pop, ADULTS, fatal(), FEM, TProtoQuanti::get_allele_model(), TProtoQuanti::get_locus_ID(), TProtoQuanti::get_num_locus(), TProtoQuanti::get_num_traits(), FileHandler::get_path(), FileHandler::get_service(), TProtoQuanti::get_type(), Metapop::getCurrentAge(), FileServices::getGenerationReplicateFileName(), GeneticMap::getGeneticMap(), Metapop::getNumAgeClasses(), Metapop::getPatch(), Metapop::getPatchNbr(), MAL, message(), OFFSPRG, OFFSx, print_PLINK_FAM(), and print_PLINK_PED().
Referenced by FHwrite().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ write_TABLE()
| void TTQuantiFH::write_TABLE | ( | ) |
References TraitFileHandler< TProtoQuanti >::_FHLinkedTrait, _output_option, FileHandler::_pop, ADLTx, fatal(), FEM, TProtoQuanti::get_allele_model(), TProtoQuanti::get_dominance_model(), TProtoQuanti::get_env_var(), FileHandler::get_filename(), TProtoQuanti::get_locus_ID(), TProtoQuanti::get_num_locus(), TProtoQuanti::get_num_traits(), Metapop::getPatch(), Metapop::getPatchNbr(), MAL, message(), OFFSx, print(), and Patch::size().
Referenced by FHwrite().
 Here is the caller graph for this function:
Here is the caller graph for this function:Member Data Documentation
◆ _has_genetic_map
|
private |
Referenced by setOutputOption(), and write_PLINK().
◆ _output_option
|
private |
Referenced by FHwrite(), setOutputOption(), and write_TABLE().
The documentation for this class was generated from the following files:






 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
