#include <ttquanti.h>
 Inheritance diagram for TTQuantiSH:
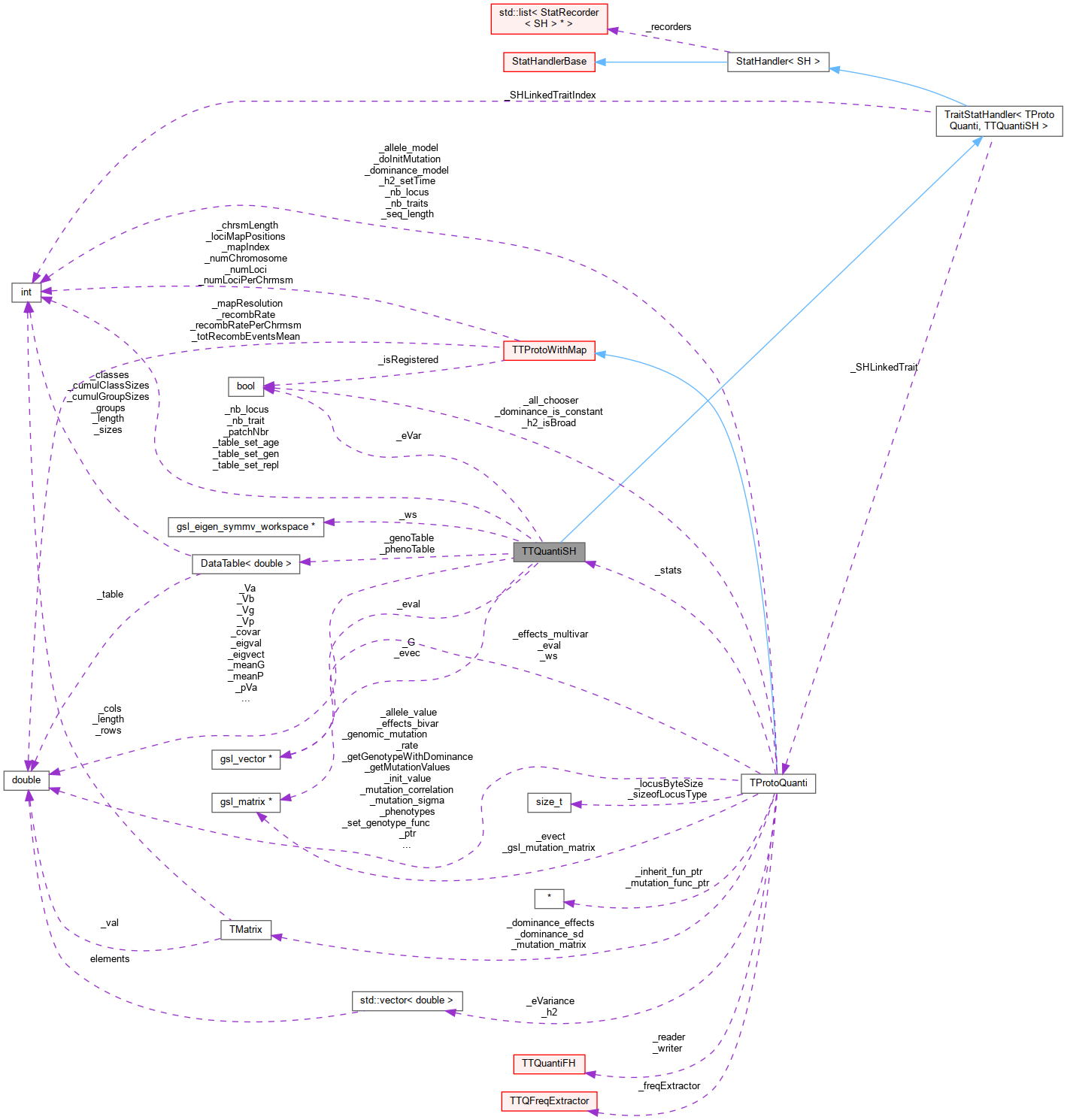
Inheritance diagram for TTQuantiSH: Collaboration diagram for TTQuantiSH:
Collaboration diagram for TTQuantiSH:Public Member Functions | |
| TTQuantiSH (TProtoQuanti *TP) | |
| virtual | ~TTQuantiSH () |
| void | resetPtrs () |
| virtual void | init () |
| virtual bool | setStatRecorders (std::string &token) |
| void | addQuanti (age_t AGE) |
| void | addQuantiPerPatch (age_t AGE) |
| void | addAvgPerPatch (age_t AGE) |
| void | addGenotPerPatch (age_t AGE) |
| void | addVarPerPatch (age_t AGE) |
| void | addCovarPerPatch (age_t AGE) |
| void | addEigenPerPatch (age_t AGE) |
| void | addEigenValuesPerPatch (age_t AGE) |
| void | addEigenVect1PerPatch (age_t AGE) |
| void | addSkewPerPatch (age_t AGE) |
| void | setDataTables (age_t AGE) |
| void | setAdultStats () |
| void | setOffsprgStats () |
| void | setStats (age_t AGE) |
| double | getMeanGenot (unsigned int i) |
| double | getMeanPhenot (unsigned int i) |
| double | getVa (unsigned int i) |
| double | getVg (unsigned int i) |
| double | getVb (unsigned int i) |
| double | getVp (unsigned int i) |
| double | getQst (unsigned int i) |
| double | getCovar (unsigned int i) |
| double | getMeanGenotPerPatch (unsigned int i, unsigned int p) |
| double | getMeanPhenotPerPatch (unsigned int i, unsigned int p) |
| double | getVaPerPatch (unsigned int i, unsigned int p) |
| double | getVpPerPatch (unsigned int i, unsigned int p) |
| double | getEigenValuePerPatch (unsigned int i, unsigned int p) |
| double | getCovarPerPatch (unsigned int p, unsigned int i) |
| double | getEigenVectorEltPerPatch (unsigned int p, unsigned int v) |
| double | getSkewPerPatch (unsigned int i, unsigned int p) |
| vector< double > | getSNPalleleFreqInPatch (Patch *patch, const age_idx AGE) |
| vector< double > | getVaWithDominance (Patch *curPop, const age_idx AGE) |
| computation of the additive genetic variance from the average excess of each allele exact under random mating only More... | |
| vector< double > | getVaNoDominance (Patch *curPop, const age_idx AGE) |
 Public Member Functions inherited from TraitStatHandler< TProtoQuanti, TTQuantiSH > Public Member Functions inherited from TraitStatHandler< TProtoQuanti, TTQuantiSH > | |
| TraitStatHandler (TProtoQuanti *trait_proto) | |
| virtual | ~TraitStatHandler () |
 Public Member Functions inherited from StatHandler< SH > Public Member Functions inherited from StatHandler< SH > | |
| StatHandler () | |
| virtual | ~StatHandler () |
| virtual void | clear () |
| Empties the _recorders list, they are destroyed in StatHandlerBase::reset(). More... | |
| virtual StatRecorder< SH > * | add (std::string Title, std::string Name, age_t AGE, unsigned int ARG1, unsigned int ARG2, double(SH::*getStatNoArg)(void), double(SH::*getStatOneArg)(unsigned int), double(SH::*getStatTwoArg)(unsigned int, unsigned int), void(SH::*setStat)(void)) |
| Adds a StatRecorder to the list, it is also added to the StatHandlerBase::_stats list. More... | |
 Public Member Functions inherited from StatHandlerBase Public Member Functions inherited from StatHandlerBase | |
| StatHandlerBase () | |
| virtual | ~StatHandlerBase () |
| virtual void | reset () |
| Empties the _stats list and calls clear() (defined in the derived class). More... | |
| Metapop * | get_pop_ptr () |
| void | set_service (StatServices *srv) |
| StatServices * | get_service () |
| unsigned int | getOccurrence () |
| unsigned int | getNumOccurrences () |
| unsigned int | getCurrentOccurrence () |
| unsigned int | getNbRecorders () |
| std::list< StatRecBase * > & | getStats () |
| virtual void | add (StatRecBase *rec) |
| virtual void | update () |
| This function is left empty as the StatServices calls StatRecorder::setVal directly. More... | |
 Public Member Functions inherited from Handler Public Member Functions inherited from Handler | |
| virtual | ~Handler () |
Private Attributes | |
| double * | _meanP |
| double * | _meanG |
| double * | _Va |
| double * | _Vg |
| double * | _Vb |
| double * | _Vp |
| double * | _covar |
| double ** | _pmeanP |
| double ** | _pmeanG |
| double ** | _pVa |
| double ** | _pVp |
| double ** | _pcovar |
| double ** | _peigval |
| double ** | _peigvect |
| bool | _eVar |
| unsigned int | _num_locus |
| unsigned int | _num_trait |
| unsigned int | _patchNbr |
| gsl_matrix * | _G |
| gsl_matrix * | _evec |
| gsl_vector * | _eval |
| gsl_eigen_symmv_workspace * | _ws |
| DataTable< double > | _phenoTable |
| DataTable< double > | _genoTable |
| unsigned int | _table_set_gen |
| unsigned int | _table_set_age |
| unsigned int | _table_set_repl |
Additional Inherited Members | |
 Protected Types inherited from StatHandler< SH > Protected Types inherited from StatHandler< SH > | |
| typedef std::list< StatRecorder< SH > * >::iterator | REC_IT |
 Protected Attributes inherited from TraitStatHandler< TProtoQuanti, TTQuantiSH > Protected Attributes inherited from TraitStatHandler< TProtoQuanti, TTQuantiSH > | |
| TProtoQuanti * | _SHLinkedTrait |
| Pointer to a TraitProtoype object. More... | |
| int | _SHLinkedTraitIndex |
| Index of the trait in the Individual::Traits table. More... | |
 Protected Attributes inherited from StatHandler< SH > Protected Attributes inherited from StatHandler< SH > | |
| std::list< StatRecorder< SH > * > | _recorders |
| The list of stat recorders. More... | |
 Protected Attributes inherited from StatHandlerBase Protected Attributes inherited from StatHandlerBase | |
| Metapop * | _pop |
| Link to the current population, set through the link to the StatService. More... | |
Detailed Description
Constructor & Destructor Documentation
◆ TTQuantiSH()
|
inline |
◆ ~TTQuantiSH()
|
inlinevirtual |
References resetPtrs().
Member Function Documentation
◆ addAvgPerPatch()
| void TTQuantiSH::addAvgPerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, getMeanPhenotPerPatch(), Metapop::getPatchNbr(), tstring::int2str(), OFFSPRG, setAdultStats(), and setOffsprgStats().
Referenced by addQuantiPerPatch(), and setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addCovarPerPatch()
| void TTQuantiSH::addCovarPerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, getCovarPerPatch(), Metapop::getPatchNbr(), tstring::int2str(), OFFSPRG, setAdultStats(), and setOffsprgStats().
Referenced by addQuantiPerPatch(), and setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addEigenPerPatch()
| void TTQuantiSH::addEigenPerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, getEigenValuePerPatch(), getEigenVectorEltPerPatch(), Metapop::getPatchNbr(), tstring::int2str(), OFFSPRG, setAdultStats(), setOffsprgStats(), and warning().
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addEigenValuesPerPatch()
| void TTQuantiSH::addEigenValuesPerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, getEigenValuePerPatch(), Metapop::getPatchNbr(), tstring::int2str(), OFFSPRG, setAdultStats(), setOffsprgStats(), and warning().
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addEigenVect1PerPatch()
| void TTQuantiSH::addEigenVect1PerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, getEigenVectorEltPerPatch(), Metapop::getPatchNbr(), tstring::int2str(), OFFSPRG, setAdultStats(), setOffsprgStats(), and warning().
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addGenotPerPatch()
| void TTQuantiSH::addGenotPerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, getMeanGenotPerPatch(), Metapop::getPatchNbr(), tstring::int2str(), OFFSPRG, setAdultStats(), and setOffsprgStats().
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addQuanti()
| void TTQuantiSH::addQuanti | ( | age_t | AGE | ) |
References _eVar, _num_trait, StatHandlerBase::_pop, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTrait, StatHandler< SH >::add(), ADULTS, ALL, TProtoQuanti::do_epistasis(), TProtoQuanti::get_dominance_model(), getCovar(), getMeanPhenot(), Metapop::getPatchNbr(), getQst(), getVa(), getVb(), getVp(), tstring::int2str(), OFFSPRG, setAdultStats(), and setOffsprgStats().
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addQuantiPerPatch()
| void TTQuantiSH::addQuantiPerPatch | ( | age_t | AGE | ) |
References addAvgPerPatch(), addCovarPerPatch(), addVarPerPatch(), ADULTS, ALL, and OFFSPRG.
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addSkewPerPatch()
| void TTQuantiSH::addSkewPerPatch | ( | age_t | AGE | ) |
References _num_trait, StatHandlerBase::_pop, StatHandler< SH >::add(), ADULTS, ALL, Metapop::getPatchNbr(), getSkewPerPatch(), tstring::int2str(), OFFSPRG, setAdultStats(), and setOffsprgStats().
Referenced by setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ addVarPerPatch()
| void TTQuantiSH::addVarPerPatch | ( | age_t | AGE | ) |
References _eVar, _num_trait, StatHandlerBase::_pop, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTrait, StatHandler< SH >::add(), ADULTS, ALL, TProtoQuanti::do_epistasis(), TProtoQuanti::get_dominance_model(), Metapop::getPatchNbr(), getVaPerPatch(), getVpPerPatch(), tstring::int2str(), OFFSPRG, setAdultStats(), and setOffsprgStats().
Referenced by addQuantiPerPatch(), and setStatRecorders().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getCovar()
|
inline |
◆ getCovarPerPatch()
|
inline |
◆ getEigenValuePerPatch()
|
inline |
References _peigval.
Referenced by addEigenPerPatch(), and addEigenValuesPerPatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getEigenVectorEltPerPatch()
|
inline |
References _peigvect.
Referenced by addEigenPerPatch(), and addEigenVect1PerPatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getMeanGenot()
◆ getMeanGenotPerPatch()
|
inline |
◆ getMeanPhenot()
|
inline |
◆ getMeanPhenotPerPatch()
|
inline |
◆ getQst()
|
inline |
◆ getSkewPerPatch()
| double TTQuantiSH::getSkewPerPatch | ( | unsigned int | i, |
| unsigned int | p | ||
| ) |
References _phenoTable, _pmeanP, DataTable< T >::getClassWithinGroup(), and DataTable< T >::size().
Referenced by addSkewPerPatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getSNPalleleFreqInPatch()
References TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTrait, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTraitIndex, fatal(), Patch::get(), TTQuanti::get_allele_bit(), TProtoQuanti::get_allele_model(), TProtoQuanti::get_seq_length(), Individual::getTrait(), and Patch::size().
Referenced by getVaNoDominance(), and getVaWithDominance().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getVa()
|
inline |
◆ getVaNoDominance()
CYCLE THROUGH ALL TRAITS
References _num_locus, _num_trait, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTrait, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTraitIndex, _Vg, FEM, Patch::get(), TMatrix::get(), TTQuanti::get_additive_genotype(), TProtoQuanti::get_diallele_values(), getSNPalleleFreqInPatch(), Individual::getTrait(), MAL, message(), my_variance_with_fixed_mean(), and Patch::size().
Referenced by LCE_QuantiModifier::setParameters(), and LCE_Breed_Quanti::setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getVaPerPatch()
|
inline |
◆ getVaWithDominance()
computation of the additive genetic variance from the average excess of each allele exact under random mating only
CYCLE THROUGH ALL TRAITS
References _num_trait, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTrait, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTraitIndex, _Vg, fatal(), Patch::get(), TMatrix::get(), TTQuanti::get_allele_bit(), TProtoQuanti::get_allele_model(), TProtoQuanti::get_diallele_values(), TTQuanti::get_full_genotype(), TProtoQuanti::get_genotype_with_dominance(), TProtoQuanti::get_h2_isBroad(), TProtoQuanti::get_locus_ID(), TProtoQuanti::get_locus_seq_pos(), TProtoQuanti::get_num_locus(), Patch::getID(), getSNPalleleFreqInPatch(), Individual::getTrait(), message(), my_variance_with_fixed_mean(), and Patch::size().
Referenced by LCE_QuantiModifier::setParameters(), and LCE_Breed_Quanti::setParameters().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getVb()
|
inline |
◆ getVg()
|
inline |
References _Vg.
Referenced by LCE_QuantiModifier::setVefromVa().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ getVp()
|
inline |
◆ getVpPerPatch()
|
inline |
◆ init()
|
virtual |
Reimplemented from StatHandlerBase.
References _covar, _eval, _eVar, _evec, _G, _meanG, _meanP, _num_locus, _num_trait, _patchNbr, _pcovar, _peigval, _peigvect, _pmeanG, _pmeanP, StatHandlerBase::_pop, _pVa, _pVp, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTrait, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTraitIndex, _Va, _Vb, _Vg, _Vp, _ws, TProtoQuanti::get_env_var(), TProtoQuanti::get_heritability(), TProtoQuanti::get_num_locus(), TProtoQuanti::get_num_traits(), TProtoQuanti::get_type(), Metapop::getPatchNbr(), IndFactory::getTraitIndex(), StatHandlerBase::init(), and resetPtrs().
◆ resetPtrs()
| void TTQuantiSH::resetPtrs | ( | ) |
◆ setAdultStats()
|
inline |
References ADULTS, and setStats().
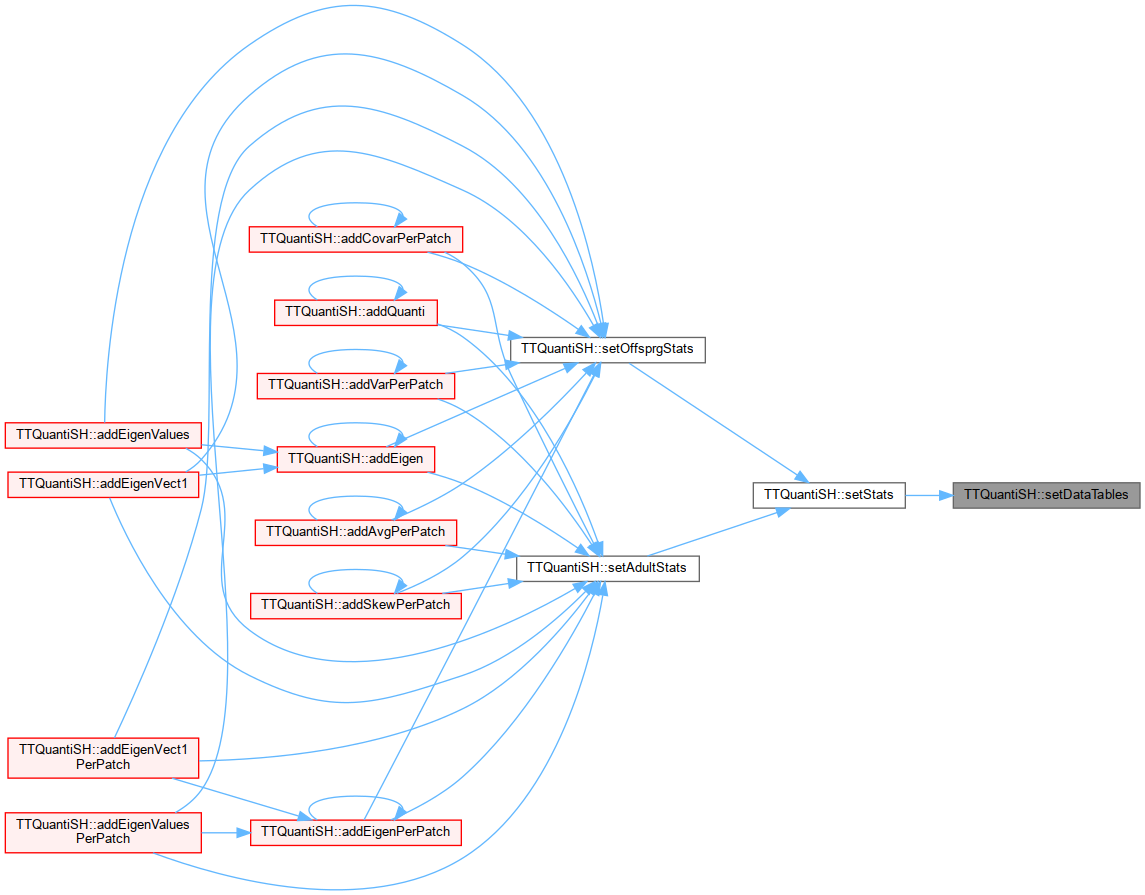
Referenced by addAvgPerPatch(), addCovarPerPatch(), addEigenPerPatch(), addEigenValuesPerPatch(), addEigenVect1PerPatch(), addGenotPerPatch(), addQuanti(), addSkewPerPatch(), and addVarPerPatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setDataTables()
| void TTQuantiSH::setDataTables | ( | age_t | AGE | ) |
References _genoTable, _num_trait, _phenoTable, StatHandlerBase::_pop, TraitStatHandler< TProtoQuanti, TTQuantiSH >::_SHLinkedTraitIndex, ADLTx, ADULTS, fatal(), FEM, Metapop::getPatch(), Metapop::getPatchNbr(), MAL, OFFSx, Metapop::size(), Patch::size(), DataTable< T >::size(), store_quanti_trait_values(), and DataTable< T >::update().
Referenced by setStats().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setOffsprgStats()
|
inline |
References OFFSPRG, and setStats().
Referenced by addAvgPerPatch(), addCovarPerPatch(), addEigenPerPatch(), addEigenValuesPerPatch(), addEigenVect1PerPatch(), addGenotPerPatch(), addQuanti(), addSkewPerPatch(), and addVarPerPatch().
 Here is the caller graph for this function:
Here is the caller graph for this function:◆ setStatRecorders()
|
virtual |
Implements StatHandlerBase.
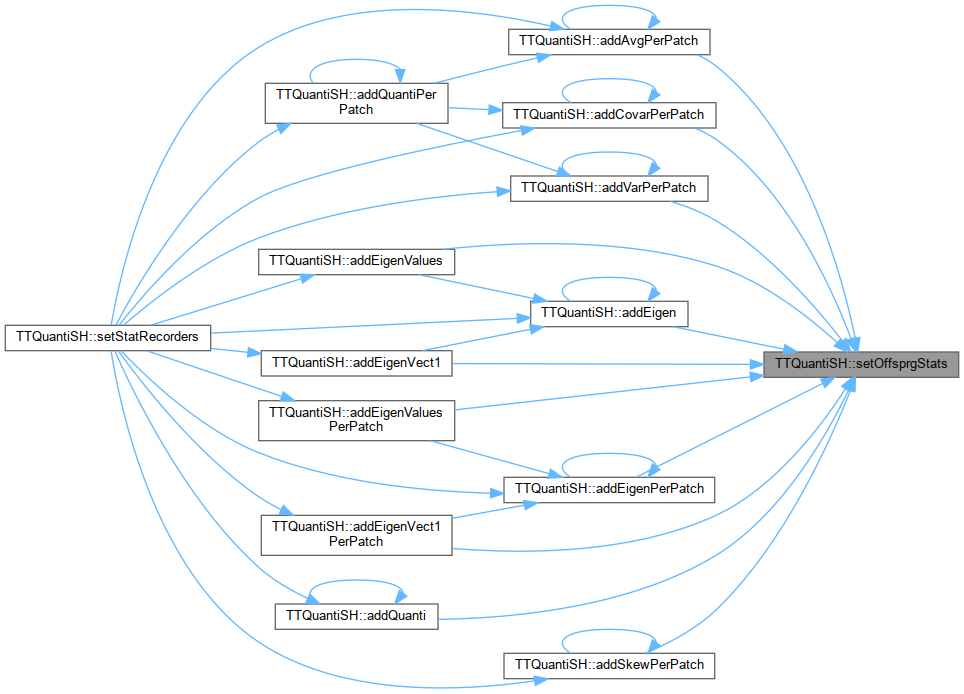
References addAvgPerPatch(), addCovarPerPatch(), addEigenPerPatch(), addEigenValuesPerPatch(), addEigenVect1PerPatch(), addGenotPerPatch(), addQuanti(), addQuantiPerPatch(), addSkewPerPatch(), addVarPerPatch(), ADULTS, ALL, message(), and OFFSPRG.
◆ setStats()
| void TTQuantiSH::setStats | ( | age_t | AGE | ) |
References _covar, _eval, _evec, _G, _genoTable, _meanG, _meanP, _num_trait, _patchNbr, _pcovar, _peigval, _peigvect, _phenoTable, _pmeanG, _pmeanP, StatHandlerBase::_pop, _pVa, _pVp, _table_set_age, _table_set_gen, _table_set_repl, _Va, _Vb, _Vp, _ws, fatal(), DataTable< T >::getClassWithinGroup(), Metapop::getCurrentGeneration(), Metapop::getCurrentReplicate(), DataTable< T >::getGroup(), Metapop::getPatchNbr(), my_covariance_no_nan(), my_mean(), my_mean_no_nan(), my_variance_with_fixed_mean(), my_variance_with_fixed_mean_no_nan(), setDataTables(), Metapop::size(), and warning().
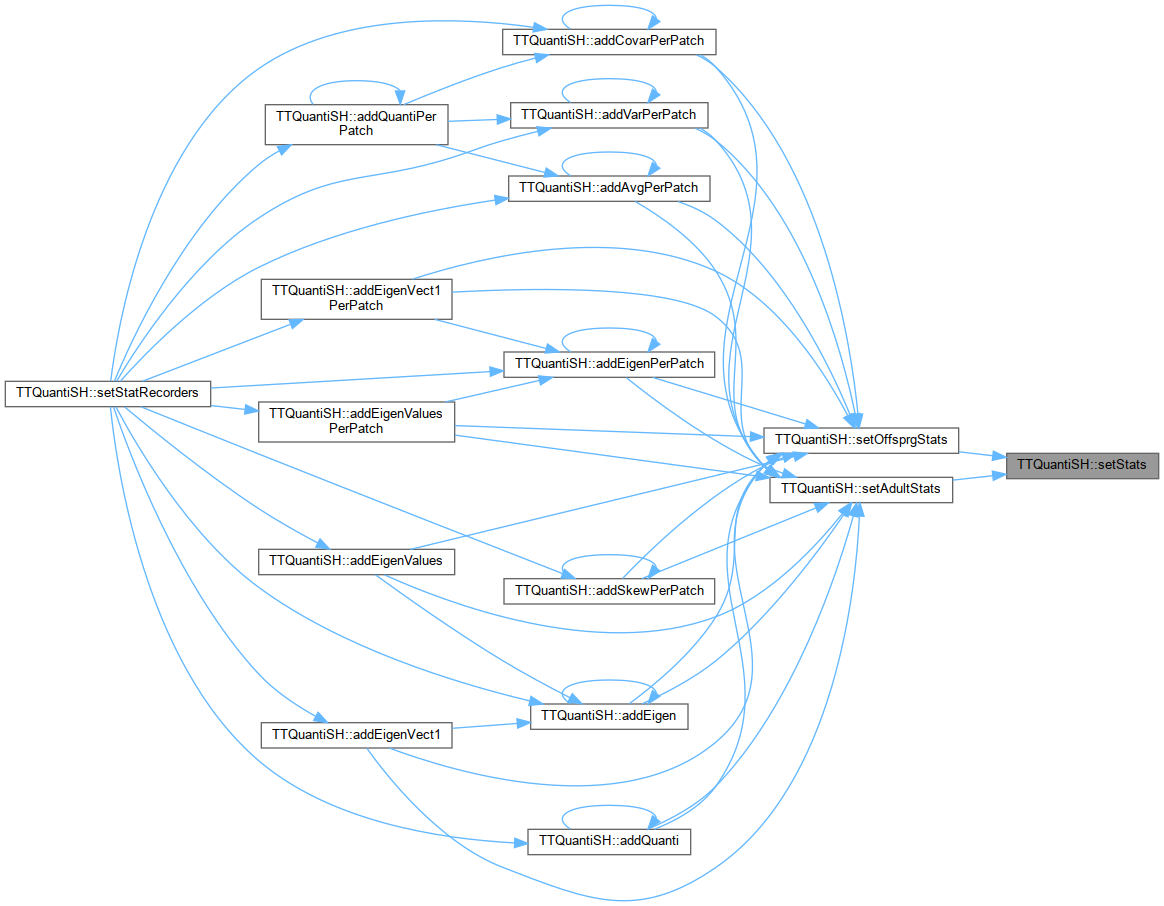
Referenced by setAdultStats(), and setOffsprgStats().
 Here is the caller graph for this function:
Here is the caller graph for this function:Member Data Documentation
◆ _covar
|
private |
Referenced by getCovar(), init(), resetPtrs(), and setStats().
◆ _eval
|
private |
Referenced by init(), resetPtrs(), and setStats().
◆ _eVar
|
private |
Referenced by addQuanti(), addVarPerPatch(), and init().
◆ _evec
|
private |
Referenced by init(), resetPtrs(), and setStats().
◆ _G
|
private |
Referenced by init(), resetPtrs(), and setStats().
◆ _genoTable
|
private |
Referenced by setDataTables(), and setStats().
◆ _meanG
|
private |
Referenced by getMeanGenot(), init(), resetPtrs(), and setStats().
◆ _meanP
|
private |
Referenced by getMeanPhenot(), init(), resetPtrs(), and setStats().
◆ _num_locus
|
private |
Referenced by getVaNoDominance(), and init().
◆ _num_trait
|
private |
◆ _patchNbr
|
private |
Referenced by init(), resetPtrs(), and setStats().
◆ _pcovar
|
private |
Referenced by getCovarPerPatch(), init(), resetPtrs(), and setStats().
◆ _peigval
|
private |
Referenced by getEigenValuePerPatch(), init(), resetPtrs(), and setStats().
◆ _peigvect
|
private |
Referenced by getEigenVectorEltPerPatch(), init(), resetPtrs(), and setStats().
◆ _phenoTable
|
private |
Referenced by getSkewPerPatch(), setDataTables(), and setStats().
◆ _pmeanG
|
private |
Referenced by getMeanGenotPerPatch(), init(), resetPtrs(), and setStats().
◆ _pmeanP
|
private |
Referenced by getMeanPhenotPerPatch(), getSkewPerPatch(), init(), resetPtrs(), and setStats().
◆ _pVa
|
private |
Referenced by getVaPerPatch(), init(), resetPtrs(), and setStats().
◆ _pVp
|
private |
Referenced by getVpPerPatch(), init(), resetPtrs(), and setStats().
◆ _table_set_age
|
private |
Referenced by setStats().
◆ _table_set_gen
|
private |
Referenced by setStats().
◆ _table_set_repl
|
private |
Referenced by setStats().
◆ _Va
|
private |
Referenced by getQst(), getVa(), init(), resetPtrs(), and setStats().
◆ _Vb
|
private |
Referenced by getQst(), getVb(), init(), resetPtrs(), and setStats().
◆ _Vg
|
private |
Referenced by getVaNoDominance(), getVaWithDominance(), getVg(), init(), and resetPtrs().
◆ _Vp
|
private |
Referenced by getVp(), init(), resetPtrs(), and setStats().
◆ _ws
|
private |
Referenced by init(), resetPtrs(), and setStats().
The documentation for this class was generated from the following files:




















 1.9.1 -- Nemo is hosted on
1.9.1 -- Nemo is hosted on 
